UniProtKB/Swiss-Prot protein knowledgebase release 2026_01 statistics
1. INTRODUCTION
Release 2026_01 of 28-Jan-2026 of UniProtKB/Swiss-Prot contains 574627 sequence
entries, curated from 310243 unique references and comprising 208482574 amino acids.
987 sequences have been added since release 2025_04, the sequence data of
180 existing entries has been updated and the annotations of
380278 entries have been revised.
Number of fragments: 9266
Number of additional sequences produced by alternative splicing, initiation or promoter usage, or ribosomal frameshifting: 41333
Protein existence (PE): entries %
1: Evidence at protein level 120182 20.9%
2: Evidence at transcript level 54408 9.5%
3: Inferred from homology 385597 67.1%
4: Predicted 12709 2.2%
5: Uncertain 1731 0.3%
The growth of the database is summarized below.
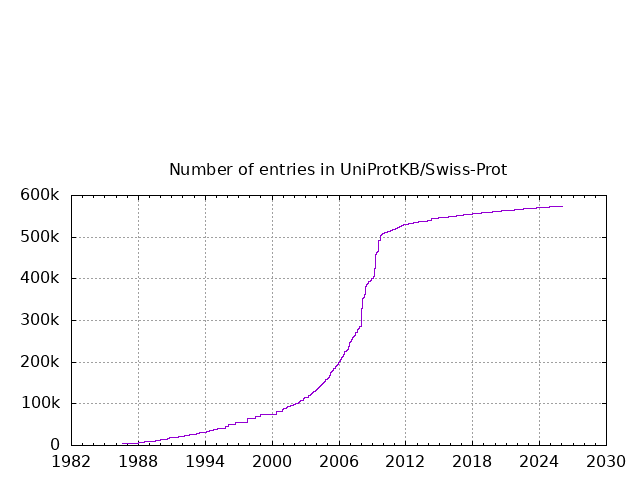 2. TAXONOMIC ORIGIN
Total number of species represented in this release of UniProtKB/Swiss-Prot: 14846
The first twenty species represent 123389 sequences: 21.5 % of the total
number of entries.
2.1 Table of the frequency of occurrence of species
Species represented 1x: 6040
2x: 2144
3x: 1167
4x: 793
5x: 549
6x: 448
7x: 332
8x: 295
9x: 238
10x: 162
11- 20x: 860
21- 50x: 521
51-100x: 235
>100x: 1062
2.2 Table of the most represented species
------ --------- --------------------------------------------
Number Frequency Species
------ --------- --------------------------------------------
1 20431 Homo sapiens (Human)
2 17252 Mus musculus (Mouse)
3 16418 Arabidopsis thaliana (Mouse-ear cress)
4 8226 Rattus norvegicus (Rat)
5 6733 Saccharomyces cerevisiae (strain ATCC 204508 / S288c) (Baker's yeast)
6 6052 Bos taurus (Bovine)
7 5129 Schizosaccharomyces pombe (strain 972 / ATCC 24843) (Fission yeast)
8 4531 Escherichia coli (strain K12)
9 4499 Caenorhabditis elegans
10 4197 Oryza sativa subsp. japonica (Rice)
11 4191 Bacillus subtilis (strain 168)
12 4163 Dictyostelium discoideum (Social amoeba)
13 3868 Drosophila melanogaster (Fruit fly)
14 3514 Xenopus laevis (African clawed frog)
15 3369 Danio rerio (Zebrafish) (Brachydanio rerio)
16 2338 Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv)
17 2314 Gallus gallus (Chicken)
18 2218 Pongo abelii (Sumatran orangutan) (Pongo pygmaeus abelii)
19 2047 Escherichia coli O157:H7
20 1899 Mycobacterium tuberculosis (strain CDC 1551 / Oshkosh)
21 1831 Salmonella typhimurium (strain LT2 / SGSC1412 / ATCC 700720)
22 1787 Methanocaldococcus jannaschii
23 1713 Xenopus tropicalis (Western clawed frog) (Silurana tropicalis)
24 1703 Haemophilus influenzae (strain ATCC 51907 / DSM 11121 / KW20 / Rd)
25 1702 Escherichia coli O6:H1 (strain CFT073 / ATCC 700928 / UPEC)
26 1696 Shigella flexneri
27 1479 Pseudomonas aeruginosa
28 1462 Sus scrofa (Pig)
29 1349 Salmonella typhi
30 1244 Mycobacterium bovis (strain ATCC BAA-935 / AF2122/97)
31 1197 Candida albicans (strain SC5314 / ATCC MYA-2876) (Yeast)
32 1176 Macaca fascicularis (Crab-eating macaque) (Cynomolgus monkey)
33 1108 Synechocystis sp. (strain ATCC 27184 / PCC 6803 / Kazusa)
34 1038 Archaeoglobus fulgidus
35 1030 Yersinia pestis
36 1028 Emericella nidulans
37 1000 Vibrio cholerae serotype O1 (strain ATCC 39315 / El Tor Inaba N16961)
38 979 Oryctolagus cuniculus (Rabbit)
39 970 Neurospora crassa
40 959 Aspergillus fumigatus (strain ATCC MYA-4609 / CBS 101355 / FGSC A1100 / Af293)
41 942 Staphylococcus aureus (strain Mu50 / ATCC 700699)
42 930 Salmonella paratyphi A (strain ATCC 9150 / SARB42)
43 929 Staphylococcus aureus (strain N315)
44 928 Eremothecium gossypii
45 920 Kluyveromyces lactis
46 909 Acanthamoeba polyphaga mimivirus (APMV)
47 905 Staphylococcus aureus (strain COL)
48 896 Staphylococcus aureus (strain MW2)
49 894 Escherichia coli O6:K15:H31 (strain 536 / UPEC)
50 892 Rhizobium meliloti (strain 1021) (Ensifer meliloti) (Sinorhizobium meliloti)
51 890 Candida glabrata
52 890 Staphylococcus aureus (strain MSSA476)
53 888 Staphylococcus aureus (strain MRSA252)
54 882 Salmonella choleraesuis (strain SC-B67)
55 879 Shigella sonnei (strain Ss046)
56 877 Oryza sativa subsp. indica (Rice)
57 863 Yersinia pseudotuberculosis serotype I (strain IP32953)
58 857 Canis lupus familiaris (Dog) (Canis familiaris)
59 850 Zea mays (Maize)
60 847 Escherichia coli O9:H4 (strain HS)
61 838 Escherichia coli O139:H28 (strain E24377A / ETEC)
62 829 Shigella boydii serotype 4 (strain Sb227)
63 825 Escherichia coli (strain UTI89 / UPEC)
64 822 Escherichia coli
65 822 Shigella dysenteriae serotype 1 (strain Sd197)
66 819 Streptomyces coelicolor (strain ATCC BAA-471 / A3(2) / M145)
67 813 Staphylococcus aureus (strain NCTC 8325 / PS 47)
68 804 Pectobacterium atrosepticum (strain SCRI 1043 / ATCC BAA-672)
69 796 Vibrio parahaemolyticus serotype O3:K6 (strain RIMD 2210633)
70 791 Escherichia coli (strain SMS-3-5 / SECEC)
71 788 Aquifex aeolicus (strain VF5)
72 779 Escherichia coli O127:H6 (strain E2348/69 / EPEC)
73 771 Escherichia coli (strain K12 / DH10B)
74 770 Pasteurella multocida (strain Pm70)
75 767 Escherichia coli O17:K52:H18 (strain UMN026 / ExPEC)
76 765 Escherichia coli (strain K12 / MC4100 / BW2952)
77 762 Escherichia coli (strain 55989 / EAEC)
78 761 Escherichia coli O8 (strain IAI1)
79 760 Shigella flexneri serotype 5b (strain 8401)
80 760 Staphylococcus epidermidis
81 760 Staphylococcus epidermidis (strain ATCC 12228 / FDA PCI 1200)
82 759 Escherichia coli O45:K1 (strain S88 / ExPEC)
83 758 Bacillus anthracis
84 756 Escherichia coli (strain SE11)
85 753 Escherichia coli O7:K1 (strain IAI39 / ExPEC)
86 749 Photorhabdus laumondii subsp. laumondii (strain DSM 15139 / CIP 105565 / TT01)
87 748 Escherichia coli O157:H7 (strain EC4115 / EHEC)
88 744 Halalkalibacterium halodurans
89 739 Yersinia enterocolitica serotype O:8 / biotype 1B (strain NCTC 13174 / 8081)
90 739 Escherichia coli
91 738 Pseudomonas putida
92 733 Vibrio vulnificus (strain CMCP6)
93 731 Escherichia coli O81 (strain ED1a)
94 722 Salmonella enteritidis PT4 (strain P125109)
95 719 Klebsiella pneumoniae subsp. pneumoniae (strain ATCC 700721 / MGH 78578)
96 718 Vibrio vulnificus (strain YJ016)
97 716 Salmonella paratyphi B (strain ATCC BAA-1250 / SPB7)
98 715 Enterobacter sp. (strain 638)
99 715 Escherichia coli O1:K1 / APEC
100 715 Yersinia pestis bv. Antiqua (strain Nepal516)
101 714 Salmonella paratyphi A (strain AKU_12601)
102 713 Yersinia pseudotuberculosis serotype O:1b (strain IP 31758)
103 713 Salmonella newport (strain SL254)
104 713 Salmonella agona (strain SL483)
105 712 Salmonella schwarzengrund (strain CVM19633)
106 711 Yersinia pestis bv. Antiqua (strain Antiqua)
107 710 Salmonella heidelberg (strain SL476)
108 708 Nostoc sp. (strain PCC 7120 / SAG 25.82 / UTEX 2576)
109 702 Salmonella dublin (strain CT_02021853)
110 699 Klebsiella variicola (strain 342) (Klebsiella pneumoniae)
111 698 Shigella boydii serotype 18 (strain CDC 3083-94 / BS512)
112 695 Escherichia fergusonii
113 692 Pan troglodytes (Chimpanzee)
114 686 Mycoplasma pneumoniae (strain ATCC 29342 / M129 / Subtype 1)
115 684 Salmonella gallinarum (strain 287/91 / NCTC 13346)
116 683 Pseudomonas syringae pv. tomato (strain ATCC BAA-871 / DC3000)
117 679 Agrobacterium fabrum (strain C58 / ATCC 33970) (Agrobacterium tumefaciens
118 679 Citrobacter koseri (strain ATCC BAA-895 / CDC 4225-83 / SGSC4696)
119 679 Staphylococcus aureus (strain USA300)
120 672 Serratia proteamaculans (strain 568)
121 672 Bacillus cereus
122 669 Mycobacterium leprae (strain TN)
123 667 Yarrowia lipolytica (strain CLIB 122 / E 150) (Yeast) (Candida lipolytica)
124 667 Bradyrhizobium diazoefficiens
125 667 Yersinia pestis (strain Pestoides F)
126 663 Shewanella oneidensis
127 658 Sinorhizobium fredii (strain NBRC 101917 / NGR234)
128 653 Debaryomyces hansenii
129 643 Staphylococcus aureus (strain bovine RF122 / ET3-1)
130 642 Yersinia pseudotuberculosis serotype O:3 (strain YPIII)
131 642 Salmonella arizonae (strain ATCC BAA-731 / CDC346-86 / RSK2980)
132 634 Yersinia pseudotuberculosis serotype IB (strain PB1/+)
133 623 Methanothermobacter thermautotrophicus
134 623 Cronobacter sakazakii (strain ATCC BAA-894) (Enterobacter sakazakii)
135 622 Treponema pallidum (strain Nichols)
136 622 Listeria monocytogenes serovar 1/2a (strain ATCC BAA-679 / EGD-e)
137 620 Pseudomonas aeruginosa (strain UCBPP-PA14)
138 615 Xanthomonas campestris pv. campestris
139 614 Mesorhizobium japonicum (Mesorhizobium loti
140 614 Staphylococcus haemolyticus (strain JCSC1435)
141 613 Helicobacter pylori (strain ATCC 700392 / 26695) (Campylobacter pylori)
142 611 Mycolicibacterium smegmatis (strain ATCC 700084 / mc(2)155)
143 605 Listeria innocua serovar 6a (strain ATCC BAA-680 / CLIP 11262)
144 604 Ralstonia nicotianae (strain ATCC BAA-1114 / GMI1000) (Ralstonia solanacearum)
145 602 Staphylococcus saprophyticus subsp. saprophyticus
146 602 Photobacterium profundum (strain SS9)
147 601 Salmonella paratyphi C (strain RKS4594)
148 600 Yersinia pestis bv. Antiqua (strain Angola)
149 595 Bacillus cereus (strain ATCC 10987 / NRS 248)
150 592 Neisseria meningitidis serogroup B (strain ATCC BAA-335 / MC58)
151 591 Pectobacterium carotovorum subsp. carotovorum (strain PC1)
152 588 Aspergillus oryzae (strain ATCC 42149 / RIB 40) (Yellow koji mold)
153 584 Rickettsia prowazekii (strain Madrid E)
154 582 Caenorhabditis briggsae
155 579 Brucella suis biovar 1 (strain 1330)
156 576 Brucella melitensis biotype 1
157 575 Caulobacter vibrioides (strain ATCC 19089 / CIP 103742 / CB 15)
158 573 Aliivibrio fischeri (strain ATCC 700601 / ES114) (Vibrio fischeri)
159 572 Buchnera aphidicola subsp. Acyrthosiphon pisum (strain APS)
160 569 Bacillus thuringiensis subsp. konkukian (strain 97-27)
161 568 Helicobacter pylori (strain J99 / ATCC 700824) (Campylobacter pylori J99)
162 568 Pseudomonas syringae pv. syringae (strain B728a)
163 567 Thermotoga maritima
164 566 Bacillus licheniformis
165 562 Bacillus cereus (strain ZK / E33L)
166 562 Buchnera aphidicola subsp. Schizaphis graminum (strain Sg)
167 561 Xanthomonas axonopodis pv. citri (strain 306)
168 559 Clostridium acetobutylicum
169 555 Pseudomonas fluorescens (strain Pf0-1)
170 554 Neisseria meningitidis serogroup A / serotype 4A (strain DSM 15465 / Z2491)
171 554 Pseudomonas fluorescens (strain ATCC BAA-477 / NRRL B-23932 / Pf-5)
172 553 Oceanobacillus iheyensis
173 547 Pseudomonas savastanoi pv. phaseolicola (Pseudomonas syringae pv. phaseolicola
174 543 Corynebacterium glutamicum
175 541 Lactococcus lactis subsp. lactis (strain IL1403) (Streptococcus lactis)
176 533 Bordetella bronchiseptica (strain ATCC BAA-588 / NCTC 13252 / RB50)
177 531 Erwinia tasmaniensis
178 530 Listeria monocytogenes serotype 4b (strain F2365)
179 529 Sodalis glossinidius (strain morsitans)
180 525 Staphylococcus aureus (strain Newman)
181 525 Deinococcus radiodurans
182 523 Vibrio cholerae serotype O1 (strain ATCC 39541 / Classical Ogawa 395 / O395)
183 522 Xylella fastidiosa (strain 9a5c)
184 519 Chromobacterium violaceum
185 519 Methanosarcina acetivorans (strain ATCC 35395 / DSM 2834 / JCM 12185 / C2A)
186 519 Streptococcus pneumoniae serotype 4 (strain ATCC BAA-334 / TIGR4)
187 516 Bordetella pertussis (strain Tohama I / ATCC BAA-589 / NCTC 13251)
188 515 Xylella fastidiosa (strain Temecula1 / ATCC 700964)
189 512 Pyrococcus furiosus (strain ATCC 43587 / DSM 3638 / JCM 8422 / Vc1)
190 512 Geobacillus kaustophilus (strain HTA426)
191 512 Haemophilus ducreyi (strain 35000HP / ATCC 700724)
192 512 Pseudomonas paraeruginosa (strain DSM 24068 / PA7) (Pseudomonas aeruginosa
193 511 Solanum lycopersicum (Tomato) (Lycopersicon esculentum)
194 511 Acinetobacter baylyi (strain ATCC 33305 / BD413 / ADP1)
195 511 Streptomyces avermitilis
196 509 Streptococcus pneumoniae (strain ATCC BAA-255 / R6)
197 508 Bordetella parapertussis (strain 12822 / ATCC BAA-587 / NCTC 13253)
198 507 Buchnera aphidicola subsp. Baizongia pistaciae (strain Bp)
199 506 Nicotiana tabacum (Common tobacco)
200 505 Saccharolobus solfataricus (strain ATCC 35092 / DSM 1617 / JCM 11322 / P2)
201 504 Pseudomonas entomophila (strain L48)
202 501 Methanosarcina mazei
203 501 Burkholderia pseudomallei (strain K96243)
204 499 Brucella abortus biovar 1 (strain 9-941)
205 499 Haemophilus influenzae (strain 86-028NP)
206 498 Thermosynechococcus vestitus (strain NIES-2133 / IAM M-273 / BP-1)
207 497 Xanthomonas campestris pv. campestris (strain 8004)
208 497 Synechococcus elongatus (strain ATCC 33912 / PCC 7942 / FACHB-805)
209 497 Pyrococcus horikoshii
210 497 Proteus mirabilis (strain HI4320)
211 496 Rickettsia conorii (strain ATCC VR-613 / Malish 7)
212 496 Shouchella clausii (strain KSM-K16) (Alkalihalobacillus clausii)
213 495 Halobacterium salinarum (strain ATCC 700922 / JCM 11081 / NRC-1)
214 493 Brucella abortus (strain 2308)
215 492 Bacillus velezensis (strain DSM 23117 / BGSC 10A6 / LMG 26770 / FZB42)
216 491 Vibrio campbellii (strain ATCC BAA-1116)
217 488 Shewanella sp. (strain MR-7)
218 486 Mannheimia succiniciproducens (strain KCTC 0769BP / MBEL55E)
219 485 Pseudomonas aeruginosa (strain LESB58)
220 485 Shewanella sp. (strain MR-4)
221 484 Staphylococcus aureus (strain Mu3 / ATCC 700698)
222 483 Mycoplasma genitalium (strain ATCC 33530 / DSM 19775 / NCTC 10195 / G37)
223 483 Lactiplantibacillus plantarum (strain ATCC BAA-793 / NCIMB 8826 / WCFS1)
224 480 Pseudomonas putida
225 478 Cupriavidus necator
226 478 Pyrococcus abyssi (strain GE5 / Orsay)
227 476 Enterococcus faecalis (strain ATCC 700802 / V583)
228 475 Burkholderia lata
229 475 Campylobacter jejuni subsp. jejuni serotype O:2
230 472 Rhodopseudomonas palustris (strain ATCC BAA-98 / CGA009)
231 471 Cereibacter sphaeroides
232 470 Clostridium perfringens (strain 13 / Type A)
233 468 Shewanella sp. (strain ANA-3)
234 468 Shewanella frigidimarina (strain NCIMB 400)
235 468 Pseudomonas putida (strain GB-1)
236 467 Aeromonas hydrophila subsp. hydrophila
237 466 Xanthomonas euvesicatoria pv. vesicatoria (strain 85-10)
238 465 Trichormus variabilis (strain ATCC 29413 / PCC 7937) (Anabaena variabilis)
239 463 Burkholderia mallei (strain ATCC 23344)
240 462 Cupriavidus pinatubonensis (strain JMP 134 / LMG 1197) (Cupriavidus necator
241 461 Ovis aries (Sheep)
242 460 Methylococcus capsulatus (strain ATCC 33009 / NCIMB 11132 / Bath)
243 457 Rickettsia felis (strain ATCC VR-1525 / URRWXCal2) (Rickettsia azadi)
244 455 Staphylococcus aureus (strain JH1)
245 455 Xanthomonas oryzae pv. oryzae (strain MAFF 311018)
246 455 Shewanella baltica (strain OS185)
247 453 Streptococcus mutans serotype c (strain ATCC 700610 / UA159)
248 453 Mycolicibacterium paratuberculosis (strain ATCC BAA-968 / K-10)
249 453 Pseudomonas putida (strain W619)
250 452 Caldanaerobacter subterraneus subsp. tengcongensis
2.3 Taxonomic distribution of the sequences
2. TAXONOMIC ORIGIN
Total number of species represented in this release of UniProtKB/Swiss-Prot: 14846
The first twenty species represent 123389 sequences: 21.5 % of the total
number of entries.
2.1 Table of the frequency of occurrence of species
Species represented 1x: 6040
2x: 2144
3x: 1167
4x: 793
5x: 549
6x: 448
7x: 332
8x: 295
9x: 238
10x: 162
11- 20x: 860
21- 50x: 521
51-100x: 235
>100x: 1062
2.2 Table of the most represented species
------ --------- --------------------------------------------
Number Frequency Species
------ --------- --------------------------------------------
1 20431 Homo sapiens (Human)
2 17252 Mus musculus (Mouse)
3 16418 Arabidopsis thaliana (Mouse-ear cress)
4 8226 Rattus norvegicus (Rat)
5 6733 Saccharomyces cerevisiae (strain ATCC 204508 / S288c) (Baker's yeast)
6 6052 Bos taurus (Bovine)
7 5129 Schizosaccharomyces pombe (strain 972 / ATCC 24843) (Fission yeast)
8 4531 Escherichia coli (strain K12)
9 4499 Caenorhabditis elegans
10 4197 Oryza sativa subsp. japonica (Rice)
11 4191 Bacillus subtilis (strain 168)
12 4163 Dictyostelium discoideum (Social amoeba)
13 3868 Drosophila melanogaster (Fruit fly)
14 3514 Xenopus laevis (African clawed frog)
15 3369 Danio rerio (Zebrafish) (Brachydanio rerio)
16 2338 Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv)
17 2314 Gallus gallus (Chicken)
18 2218 Pongo abelii (Sumatran orangutan) (Pongo pygmaeus abelii)
19 2047 Escherichia coli O157:H7
20 1899 Mycobacterium tuberculosis (strain CDC 1551 / Oshkosh)
21 1831 Salmonella typhimurium (strain LT2 / SGSC1412 / ATCC 700720)
22 1787 Methanocaldococcus jannaschii
23 1713 Xenopus tropicalis (Western clawed frog) (Silurana tropicalis)
24 1703 Haemophilus influenzae (strain ATCC 51907 / DSM 11121 / KW20 / Rd)
25 1702 Escherichia coli O6:H1 (strain CFT073 / ATCC 700928 / UPEC)
26 1696 Shigella flexneri
27 1479 Pseudomonas aeruginosa
28 1462 Sus scrofa (Pig)
29 1349 Salmonella typhi
30 1244 Mycobacterium bovis (strain ATCC BAA-935 / AF2122/97)
31 1197 Candida albicans (strain SC5314 / ATCC MYA-2876) (Yeast)
32 1176 Macaca fascicularis (Crab-eating macaque) (Cynomolgus monkey)
33 1108 Synechocystis sp. (strain ATCC 27184 / PCC 6803 / Kazusa)
34 1038 Archaeoglobus fulgidus
35 1030 Yersinia pestis
36 1028 Emericella nidulans
37 1000 Vibrio cholerae serotype O1 (strain ATCC 39315 / El Tor Inaba N16961)
38 979 Oryctolagus cuniculus (Rabbit)
39 970 Neurospora crassa
40 959 Aspergillus fumigatus (strain ATCC MYA-4609 / CBS 101355 / FGSC A1100 / Af293)
41 942 Staphylococcus aureus (strain Mu50 / ATCC 700699)
42 930 Salmonella paratyphi A (strain ATCC 9150 / SARB42)
43 929 Staphylococcus aureus (strain N315)
44 928 Eremothecium gossypii
45 920 Kluyveromyces lactis
46 909 Acanthamoeba polyphaga mimivirus (APMV)
47 905 Staphylococcus aureus (strain COL)
48 896 Staphylococcus aureus (strain MW2)
49 894 Escherichia coli O6:K15:H31 (strain 536 / UPEC)
50 892 Rhizobium meliloti (strain 1021) (Ensifer meliloti) (Sinorhizobium meliloti)
51 890 Candida glabrata
52 890 Staphylococcus aureus (strain MSSA476)
53 888 Staphylococcus aureus (strain MRSA252)
54 882 Salmonella choleraesuis (strain SC-B67)
55 879 Shigella sonnei (strain Ss046)
56 877 Oryza sativa subsp. indica (Rice)
57 863 Yersinia pseudotuberculosis serotype I (strain IP32953)
58 857 Canis lupus familiaris (Dog) (Canis familiaris)
59 850 Zea mays (Maize)
60 847 Escherichia coli O9:H4 (strain HS)
61 838 Escherichia coli O139:H28 (strain E24377A / ETEC)
62 829 Shigella boydii serotype 4 (strain Sb227)
63 825 Escherichia coli (strain UTI89 / UPEC)
64 822 Escherichia coli
65 822 Shigella dysenteriae serotype 1 (strain Sd197)
66 819 Streptomyces coelicolor (strain ATCC BAA-471 / A3(2) / M145)
67 813 Staphylococcus aureus (strain NCTC 8325 / PS 47)
68 804 Pectobacterium atrosepticum (strain SCRI 1043 / ATCC BAA-672)
69 796 Vibrio parahaemolyticus serotype O3:K6 (strain RIMD 2210633)
70 791 Escherichia coli (strain SMS-3-5 / SECEC)
71 788 Aquifex aeolicus (strain VF5)
72 779 Escherichia coli O127:H6 (strain E2348/69 / EPEC)
73 771 Escherichia coli (strain K12 / DH10B)
74 770 Pasteurella multocida (strain Pm70)
75 767 Escherichia coli O17:K52:H18 (strain UMN026 / ExPEC)
76 765 Escherichia coli (strain K12 / MC4100 / BW2952)
77 762 Escherichia coli (strain 55989 / EAEC)
78 761 Escherichia coli O8 (strain IAI1)
79 760 Shigella flexneri serotype 5b (strain 8401)
80 760 Staphylococcus epidermidis
81 760 Staphylococcus epidermidis (strain ATCC 12228 / FDA PCI 1200)
82 759 Escherichia coli O45:K1 (strain S88 / ExPEC)
83 758 Bacillus anthracis
84 756 Escherichia coli (strain SE11)
85 753 Escherichia coli O7:K1 (strain IAI39 / ExPEC)
86 749 Photorhabdus laumondii subsp. laumondii (strain DSM 15139 / CIP 105565 / TT01)
87 748 Escherichia coli O157:H7 (strain EC4115 / EHEC)
88 744 Halalkalibacterium halodurans
89 739 Yersinia enterocolitica serotype O:8 / biotype 1B (strain NCTC 13174 / 8081)
90 739 Escherichia coli
91 738 Pseudomonas putida
92 733 Vibrio vulnificus (strain CMCP6)
93 731 Escherichia coli O81 (strain ED1a)
94 722 Salmonella enteritidis PT4 (strain P125109)
95 719 Klebsiella pneumoniae subsp. pneumoniae (strain ATCC 700721 / MGH 78578)
96 718 Vibrio vulnificus (strain YJ016)
97 716 Salmonella paratyphi B (strain ATCC BAA-1250 / SPB7)
98 715 Enterobacter sp. (strain 638)
99 715 Escherichia coli O1:K1 / APEC
100 715 Yersinia pestis bv. Antiqua (strain Nepal516)
101 714 Salmonella paratyphi A (strain AKU_12601)
102 713 Yersinia pseudotuberculosis serotype O:1b (strain IP 31758)
103 713 Salmonella newport (strain SL254)
104 713 Salmonella agona (strain SL483)
105 712 Salmonella schwarzengrund (strain CVM19633)
106 711 Yersinia pestis bv. Antiqua (strain Antiqua)
107 710 Salmonella heidelberg (strain SL476)
108 708 Nostoc sp. (strain PCC 7120 / SAG 25.82 / UTEX 2576)
109 702 Salmonella dublin (strain CT_02021853)
110 699 Klebsiella variicola (strain 342) (Klebsiella pneumoniae)
111 698 Shigella boydii serotype 18 (strain CDC 3083-94 / BS512)
112 695 Escherichia fergusonii
113 692 Pan troglodytes (Chimpanzee)
114 686 Mycoplasma pneumoniae (strain ATCC 29342 / M129 / Subtype 1)
115 684 Salmonella gallinarum (strain 287/91 / NCTC 13346)
116 683 Pseudomonas syringae pv. tomato (strain ATCC BAA-871 / DC3000)
117 679 Agrobacterium fabrum (strain C58 / ATCC 33970) (Agrobacterium tumefaciens
118 679 Citrobacter koseri (strain ATCC BAA-895 / CDC 4225-83 / SGSC4696)
119 679 Staphylococcus aureus (strain USA300)
120 672 Serratia proteamaculans (strain 568)
121 672 Bacillus cereus
122 669 Mycobacterium leprae (strain TN)
123 667 Yarrowia lipolytica (strain CLIB 122 / E 150) (Yeast) (Candida lipolytica)
124 667 Bradyrhizobium diazoefficiens
125 667 Yersinia pestis (strain Pestoides F)
126 663 Shewanella oneidensis
127 658 Sinorhizobium fredii (strain NBRC 101917 / NGR234)
128 653 Debaryomyces hansenii
129 643 Staphylococcus aureus (strain bovine RF122 / ET3-1)
130 642 Yersinia pseudotuberculosis serotype O:3 (strain YPIII)
131 642 Salmonella arizonae (strain ATCC BAA-731 / CDC346-86 / RSK2980)
132 634 Yersinia pseudotuberculosis serotype IB (strain PB1/+)
133 623 Methanothermobacter thermautotrophicus
134 623 Cronobacter sakazakii (strain ATCC BAA-894) (Enterobacter sakazakii)
135 622 Treponema pallidum (strain Nichols)
136 622 Listeria monocytogenes serovar 1/2a (strain ATCC BAA-679 / EGD-e)
137 620 Pseudomonas aeruginosa (strain UCBPP-PA14)
138 615 Xanthomonas campestris pv. campestris
139 614 Mesorhizobium japonicum (Mesorhizobium loti
140 614 Staphylococcus haemolyticus (strain JCSC1435)
141 613 Helicobacter pylori (strain ATCC 700392 / 26695) (Campylobacter pylori)
142 611 Mycolicibacterium smegmatis (strain ATCC 700084 / mc(2)155)
143 605 Listeria innocua serovar 6a (strain ATCC BAA-680 / CLIP 11262)
144 604 Ralstonia nicotianae (strain ATCC BAA-1114 / GMI1000) (Ralstonia solanacearum)
145 602 Staphylococcus saprophyticus subsp. saprophyticus
146 602 Photobacterium profundum (strain SS9)
147 601 Salmonella paratyphi C (strain RKS4594)
148 600 Yersinia pestis bv. Antiqua (strain Angola)
149 595 Bacillus cereus (strain ATCC 10987 / NRS 248)
150 592 Neisseria meningitidis serogroup B (strain ATCC BAA-335 / MC58)
151 591 Pectobacterium carotovorum subsp. carotovorum (strain PC1)
152 588 Aspergillus oryzae (strain ATCC 42149 / RIB 40) (Yellow koji mold)
153 584 Rickettsia prowazekii (strain Madrid E)
154 582 Caenorhabditis briggsae
155 579 Brucella suis biovar 1 (strain 1330)
156 576 Brucella melitensis biotype 1
157 575 Caulobacter vibrioides (strain ATCC 19089 / CIP 103742 / CB 15)
158 573 Aliivibrio fischeri (strain ATCC 700601 / ES114) (Vibrio fischeri)
159 572 Buchnera aphidicola subsp. Acyrthosiphon pisum (strain APS)
160 569 Bacillus thuringiensis subsp. konkukian (strain 97-27)
161 568 Helicobacter pylori (strain J99 / ATCC 700824) (Campylobacter pylori J99)
162 568 Pseudomonas syringae pv. syringae (strain B728a)
163 567 Thermotoga maritima
164 566 Bacillus licheniformis
165 562 Bacillus cereus (strain ZK / E33L)
166 562 Buchnera aphidicola subsp. Schizaphis graminum (strain Sg)
167 561 Xanthomonas axonopodis pv. citri (strain 306)
168 559 Clostridium acetobutylicum
169 555 Pseudomonas fluorescens (strain Pf0-1)
170 554 Neisseria meningitidis serogroup A / serotype 4A (strain DSM 15465 / Z2491)
171 554 Pseudomonas fluorescens (strain ATCC BAA-477 / NRRL B-23932 / Pf-5)
172 553 Oceanobacillus iheyensis
173 547 Pseudomonas savastanoi pv. phaseolicola (Pseudomonas syringae pv. phaseolicola
174 543 Corynebacterium glutamicum
175 541 Lactococcus lactis subsp. lactis (strain IL1403) (Streptococcus lactis)
176 533 Bordetella bronchiseptica (strain ATCC BAA-588 / NCTC 13252 / RB50)
177 531 Erwinia tasmaniensis
178 530 Listeria monocytogenes serotype 4b (strain F2365)
179 529 Sodalis glossinidius (strain morsitans)
180 525 Staphylococcus aureus (strain Newman)
181 525 Deinococcus radiodurans
182 523 Vibrio cholerae serotype O1 (strain ATCC 39541 / Classical Ogawa 395 / O395)
183 522 Xylella fastidiosa (strain 9a5c)
184 519 Chromobacterium violaceum
185 519 Methanosarcina acetivorans (strain ATCC 35395 / DSM 2834 / JCM 12185 / C2A)
186 519 Streptococcus pneumoniae serotype 4 (strain ATCC BAA-334 / TIGR4)
187 516 Bordetella pertussis (strain Tohama I / ATCC BAA-589 / NCTC 13251)
188 515 Xylella fastidiosa (strain Temecula1 / ATCC 700964)
189 512 Pyrococcus furiosus (strain ATCC 43587 / DSM 3638 / JCM 8422 / Vc1)
190 512 Geobacillus kaustophilus (strain HTA426)
191 512 Haemophilus ducreyi (strain 35000HP / ATCC 700724)
192 512 Pseudomonas paraeruginosa (strain DSM 24068 / PA7) (Pseudomonas aeruginosa
193 511 Solanum lycopersicum (Tomato) (Lycopersicon esculentum)
194 511 Acinetobacter baylyi (strain ATCC 33305 / BD413 / ADP1)
195 511 Streptomyces avermitilis
196 509 Streptococcus pneumoniae (strain ATCC BAA-255 / R6)
197 508 Bordetella parapertussis (strain 12822 / ATCC BAA-587 / NCTC 13253)
198 507 Buchnera aphidicola subsp. Baizongia pistaciae (strain Bp)
199 506 Nicotiana tabacum (Common tobacco)
200 505 Saccharolobus solfataricus (strain ATCC 35092 / DSM 1617 / JCM 11322 / P2)
201 504 Pseudomonas entomophila (strain L48)
202 501 Methanosarcina mazei
203 501 Burkholderia pseudomallei (strain K96243)
204 499 Brucella abortus biovar 1 (strain 9-941)
205 499 Haemophilus influenzae (strain 86-028NP)
206 498 Thermosynechococcus vestitus (strain NIES-2133 / IAM M-273 / BP-1)
207 497 Xanthomonas campestris pv. campestris (strain 8004)
208 497 Synechococcus elongatus (strain ATCC 33912 / PCC 7942 / FACHB-805)
209 497 Pyrococcus horikoshii
210 497 Proteus mirabilis (strain HI4320)
211 496 Rickettsia conorii (strain ATCC VR-613 / Malish 7)
212 496 Shouchella clausii (strain KSM-K16) (Alkalihalobacillus clausii)
213 495 Halobacterium salinarum (strain ATCC 700922 / JCM 11081 / NRC-1)
214 493 Brucella abortus (strain 2308)
215 492 Bacillus velezensis (strain DSM 23117 / BGSC 10A6 / LMG 26770 / FZB42)
216 491 Vibrio campbellii (strain ATCC BAA-1116)
217 488 Shewanella sp. (strain MR-7)
218 486 Mannheimia succiniciproducens (strain KCTC 0769BP / MBEL55E)
219 485 Pseudomonas aeruginosa (strain LESB58)
220 485 Shewanella sp. (strain MR-4)
221 484 Staphylococcus aureus (strain Mu3 / ATCC 700698)
222 483 Mycoplasma genitalium (strain ATCC 33530 / DSM 19775 / NCTC 10195 / G37)
223 483 Lactiplantibacillus plantarum (strain ATCC BAA-793 / NCIMB 8826 / WCFS1)
224 480 Pseudomonas putida
225 478 Cupriavidus necator
226 478 Pyrococcus abyssi (strain GE5 / Orsay)
227 476 Enterococcus faecalis (strain ATCC 700802 / V583)
228 475 Burkholderia lata
229 475 Campylobacter jejuni subsp. jejuni serotype O:2
230 472 Rhodopseudomonas palustris (strain ATCC BAA-98 / CGA009)
231 471 Cereibacter sphaeroides
232 470 Clostridium perfringens (strain 13 / Type A)
233 468 Shewanella sp. (strain ANA-3)
234 468 Shewanella frigidimarina (strain NCIMB 400)
235 468 Pseudomonas putida (strain GB-1)
236 467 Aeromonas hydrophila subsp. hydrophila
237 466 Xanthomonas euvesicatoria pv. vesicatoria (strain 85-10)
238 465 Trichormus variabilis (strain ATCC 29413 / PCC 7937) (Anabaena variabilis)
239 463 Burkholderia mallei (strain ATCC 23344)
240 462 Cupriavidus pinatubonensis (strain JMP 134 / LMG 1197) (Cupriavidus necator
241 461 Ovis aries (Sheep)
242 460 Methylococcus capsulatus (strain ATCC 33009 / NCIMB 11132 / Bath)
243 457 Rickettsia felis (strain ATCC VR-1525 / URRWXCal2) (Rickettsia azadi)
244 455 Staphylococcus aureus (strain JH1)
245 455 Xanthomonas oryzae pv. oryzae (strain MAFF 311018)
246 455 Shewanella baltica (strain OS185)
247 453 Streptococcus mutans serotype c (strain ATCC 700610 / UA159)
248 453 Mycolicibacterium paratuberculosis (strain ATCC BAA-968 / K-10)
249 453 Pseudomonas putida (strain W619)
250 452 Caldanaerobacter subterraneus subsp. tengcongensis
2.3 Taxonomic distribution of the sequences
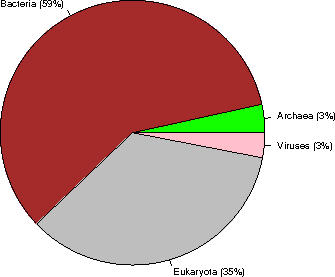 Kingdom sequences (% of the database)
Archaea 19842 ( 3%)
Bacteria 336999 ( 59%)
Eukaryota 200289 ( 35%)
Viruses 17497 ( 3%)
Within Eukaryota:
Kingdom sequences (% of the database)
Archaea 19842 ( 3%)
Bacteria 336999 ( 59%)
Eukaryota 200289 ( 35%)
Viruses 17497 ( 3%)
Within Eukaryota:
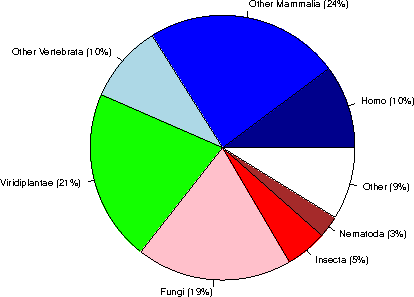 Category sequences (% of Eukaryota) (% of the complete database)
Human 20432 ( 10%) ( 4%)
Other Mammalia 47531 ( 24%) ( 8%)
Other Vertebrata 19057 ( 10%) ( 3%)
Viridiplantae 42002 ( 21%) ( 7%)
Fungi 38077 ( 19%) ( 7%)
Insecta 10104 ( 5%) ( 2%)
Nematoda 5418 ( 3%) ( 1%)
Other 17668 ( 9%) ( 3%)
3. SEQUENCE SIZE
Repartition of the sequences by size (excluding fragments)
From To Number From To Number
1- 50 10063 1001-1100 4181
51- 100 43847 1101-1200 2942
101- 150 60138 1201-1300 2236
151- 200 59834 1301-1400 2098
201- 250 58732 1401-1500 1707
251- 300 52750 1501-1600 849
301- 350 53196 1601-1700 653
351- 400 46288 1701-1800 606
401- 450 37946 1801-1900 543
451- 500 30836 1901-2000 406
501- 550 22592 2001-2100 283
551- 600 15985 2101-2200 399
601- 650 13259 2201-2300 348
651- 700 9486 2301-2400 243
701- 750 7942 2401-2500 202
751- 800 5755 >2500 1527
801- 850 4942
851- 900 5350
901- 950 4159
951-1000 3038
Category sequences (% of Eukaryota) (% of the complete database)
Human 20432 ( 10%) ( 4%)
Other Mammalia 47531 ( 24%) ( 8%)
Other Vertebrata 19057 ( 10%) ( 3%)
Viridiplantae 42002 ( 21%) ( 7%)
Fungi 38077 ( 19%) ( 7%)
Insecta 10104 ( 5%) ( 2%)
Nematoda 5418 ( 3%) ( 1%)
Other 17668 ( 9%) ( 3%)
3. SEQUENCE SIZE
Repartition of the sequences by size (excluding fragments)
From To Number From To Number
1- 50 10063 1001-1100 4181
51- 100 43847 1101-1200 2942
101- 150 60138 1201-1300 2236
151- 200 59834 1301-1400 2098
201- 250 58732 1401-1500 1707
251- 300 52750 1501-1600 849
301- 350 53196 1601-1700 653
351- 400 46288 1701-1800 606
401- 450 37946 1801-1900 543
451- 500 30836 1901-2000 406
501- 550 22592 2001-2100 283
551- 600 15985 2101-2200 399
601- 650 13259 2201-2300 348
651- 700 9486 2301-2400 243
701- 750 7942 2401-2500 202
751- 800 5755 >2500 1527
801- 850 4942
851- 900 5350
901- 950 4159
951-1000 3038
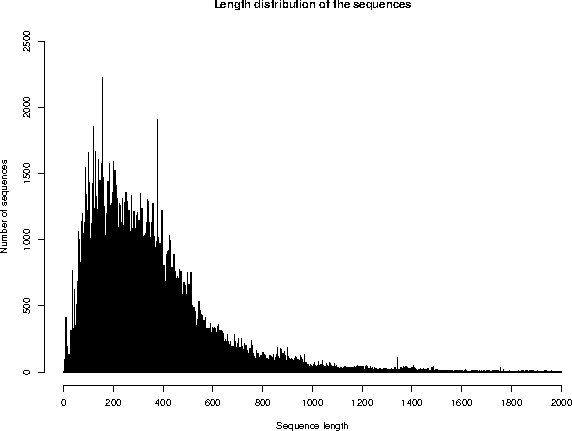 The average sequence length in UniProtKB/Swiss-Prot is 362 amino acids.
The shortest sequence is GWA_SEPOF (P83570): 2 amino acids.
The longest sequence is TITIN_MOUSE (A2ASS6): 35213 amino acids.
4. JOURNAL CITATIONS
Note: the following citation statistics reflect the number of distinct
journal citations.
Total number of journals cited in this release of UniProtKB/Swiss-Prot: 3256
4.1 Table of the frequency of journal citations
Journals cited 1x: 1018
2x: 432
3x: 233
4x: 154
5x: 128
6x: 102
7x: 64
8x: 70
9x: 63
10x: 36
11- 20x: 257
21- 50x: 283
51-100x: 144
>100x: 272
4.2 List of the most cited journals in UniProtKB/Swiss-Prot
Nb Citations Journal name
-- --------- -------------------------------------------------------------
1 27952 Journal of Biological Chemistry
2 13174 Proceedings of the National Academy of Sciences of the U.S.A.
3 7396 Journal of Bacteriology
4 6197 Biochemical and Biophysical Research Communications
5 6014 Biochemistry
6 5511 Nucleic Acids Research
7 5415 Nature
8 5172 FEBS Letters
9 5115 The EMBO Journal
10 4911 Gene
11 4725 Journal of Molecular Biology
12 4667 Molecular and Cellular Biology
13 4121 Biochimica et Biophysica Acta
14 3985 Cell
15 3712 Journal of Virology
16 3538 European Journal of Biochemistry
17 3513 Science
18 3251 Biochemical Journal
19 3002 Molecular Microbiology
20 2844 Plant Physiology
21 2829 PLoS ONE
22 2549 Genomics
23 2501 The American Journal of Human Genetics
24 2432 Journal of Cell Biology
25 2217 The Plant Cell
26 2079 Human Molecular Genetics
27 2049 The Plant Journal
28 1994 Genes and Development
29 1976 Molecular Cell
30 1947 Virology
31 1929 Plant Molecular Biology
32 1875 Nature Genetics
33 1871 Molecular Biology of the Cell
34 1859 Development
35 1774 Journal of Immunology
36 1739 Nature Communications
37 1696 Human Mutation
38 1578 Oncogene
39 1519 Structure
40 1452 Journal of Biochemistry
41 1452 Genetics
42 1444 Molecular and General Genetics
43 1440 Journal of Cell Science
44 1327 Blood
45 1302 Infection and Immunity
46 1220 Microbiology
47 1212 Developmental Biology
48 1200 Journal of General Virology
49 1179 Current Biology
50 1172 Archives of Biochemistry and Biophysics
51 1099 Scientific Reports
52 1078 Journal of Neuroscience
53 1069 Applied and Environmental Microbiology
54 1013 Acta Crystallographica, Section D
55 973 PLoS Genetics
56 963 FEMS Microbiology Letters
57 940 Cancer Research
58 912 American Journal of Physiology
59 905 Toxicon
60 903 Protein Science
61 892 Journal of Clinical Investigation
62 860 Yeast
63 856 Neuron
64 807 The Journal of Experimental Medicine
65 783 PLoS Pathogens
66 779 Human Genetics
67 776 Plant and Cell Physiology
68 775 Nature Structural and Molecular Biology
69 745 Journal of Medical Genetics
70 740 The FEBS Journal
71 715 Proteins
72 684 Nature Cell Biology
73 682 Mechanisms of Development
74 661 Bioscience, Biotechnology, and Biochemistry
75 656 Nature Structural Biology
76 649 Antimicrobial Agents and Chemotherapy
77 618 Cell Reports
78 617 Developmental Cell
79 611 Current Genetics
80 584 Journal of Neurochemistry
81 564 Journal of the American Chemical Society
82 561 Molecular Endocrinology
83 558 The Journal of Clinical Endocrinology and Metabolism
84 555 Endocrinology
85 549 Molecular and Biochemical Parasitology
86 543
87 520 Eukaryotic Cell
88 505 Experimental Cell Research
89 504 EMBO Reports
90 499 RNA
91 495 Mammalian Genome
92 490 American Journal of Medical Genetics. Part A
93 486 The FASEB Journal
94 483 Peptides
95 477 Journal of Experimental Botany
96 462 Planta
97 449 Molecular Pharmacology
98 445 Acta Crystallographica, Section F
99 444 European Journal of Human Genetics
100 437 Clinical Genetics
101 435 Immunogenetics
102 432 Molecular Plant-Microbe Interactions
103 426 Immunity
104 426 Molecular Biology and Evolution
105 422 Journal of Investigative Dermatology
106 407 Journal of Molecular Evolution
107 402 Neurology
108 400 Biochimie
109 398 DNA and Cell Biology
110 387 Biology of Reproduction
111 381 Comparative Biochemistry and Physiology
112 381 DNA Sequence
113 376 PLoS Biology
114 372 Genes to Cells
115 368 Applied Microbiology and Biotechnology
116 366 Nature Immunology
117 366 Virus Research
118 359 Journal of Lipid Research
119 356 Journal of Medicinal Chemistry
120 352 BMC Genomics
121 350 Developmental Dynamics
122 346 The New England Journal of Medicine
123 343 Brain Research. Molecular Brain Research
124 341 Annals of Neurology
125 329 European Journal of Immunology
126 321 Journal of Human Genetics
127 321 Genome Research
128 315 Nature Chemical Biology
129 315 Investigative Ophthalmology and Visual Science
130 301 Brain
131 300 Glycobiology
132 299 Biological Chemistry Hoppe-Seyler
133 286 Fungal Genetics and Biology
134 285 Journal of General Microbiology
135 285 Archives of Microbiology
136 281 Cytogenetics and Cell Genetics
137 271 Traffic
138 271 Protein Expression and Purification
139 270 Cell Research
140 269 Molecular Genetics and Metabolism
141 266 Nature Medicine
142 263 Molecular Immunology
143 263 Phytochemistry
144 258 Journal of Cellular Biochemistry
145 254 Cell Cycle
146 246 Circulation Research
147 240 Insect Biochemistry and Molecular Biology
148 240 Diabetes
149 238 New Phytologist
150 237 Chemistry and Biology
5. STATISTICS FOR SOME LINE TYPES
The following table summarizes the total number of some UniProtKB/Swiss-Prot lines,
as well as the number of entries with at least one such line, and the
frequency of the lines.
Total Number of Average
Line type / subtype number entries per entry
------------------------------------ -------- --------- ---------
References (RL) 1338719 2.33
Journal 1167917 480978 2.03 1
Submitted to EMBL/GenBank/DDBJ 159128 143141 0.28 2
Submitted to other databases 7944 7239 0.01 3
Book citation 1877 1854 <0.01 4
Plant Gene Register 613 600 <0.01 5
Unpublished observations 543 539 <0.01 6
Thesis 477 474 <0.01 7
Patent 214 207 <0.01 8
Worm Breeder's Gazette 6 6 <0.01 9
Total number of distinct authors cited in UniProtKB/Swiss-Prot: 489697
Total Number of Average
Line type / subtype number entries per entry Rank
------------------------------------ -------- --------- --------- ----
Comments (CC) 2802336 4.88
ACTIVITY REGULATION 19539 19400 0.03 17
ALLERGEN 961 961 <0.01 26
ALTERNATIVE PRODUCTS 26020 26020 0.05 14
BIOPHYSICOCHEMICAL PROPERTIES 12354 12292 0.02 20
BIOTECHNOLOGY 2342 2281 <0.01 24
CATALYTIC ACTIVITY 357782 260363 0.62 4
CAUTION 14627 14324 0.03 19
COFACTOR 135473 123006 0.24 7
DEVELOPMENTAL STAGE 14912 14783 0.03 18
DISEASE 8698 5835 0.02 21
DISRUPTION PHENOTYPE 23450 23377 0.04 16
DOMAIN 62834 53157 0.11 9
FUNCTION 500305 473996 0.87 2
INDUCTION 27070 26950 0.05 13
INTERACTION 25638 25638 0.04 15
MASS SPECTROMETRY 7707 5971 0.01 22
MISCELLANEOUS 46679 41012 0.08 11
PATHWAY 145085 131041 0.25 6
PHARMACEUTICAL 169 162 <0.01 29
POLYMORPHISM 1515 1392 <0.01 25
PTM 67522 47537 0.12 8
RNA EDITING 646 646 <0.01 28
SEQUENCE CAUTION 45415 45345 0.08 12
SIMILARITY 522913 518545 0.91 1
SUBCELLULAR LOCATION 370136 361146 0.64 3
SUBUNIT 304021 298021 0.53 5
TISSUE SPECIFICITY 52275 51527 0.09 10
TOXIC DOSE 893 714 <0.01 27
WEB RESOURCE 5355 4823 0.01 23
Total number of comment topics: 29
Total Number of Average
Line type / subtype number entries per entry Rank
------------------------------------ -------- --------- --------- ----
Features (FT) 5659423 9.85
ACT_SITE 179698 107127 0.31 10
BINDING 1287551 222598 2.24 1
CARBOHYD 126559 32187 0.22 14
CHAIN 583134 566838 1.01 2
COILED 22877 15826 0.04 25
COMPBIAS 267854 93109 0.47 8
CONFLICT 140100 48791 0.24 13
CROSSLNK 25906 9283 0.05 24
DISULFID 141462 37439 0.25 12
DNA_BIND 12308 11027 0.02 31
DOMAIN 220216 134721 0.38 9
HELIX 374823 31767 0.65 5
INIT_MET 17661 17607 0.03 26
INTRAMEM 3194 1527 0.01 34
LIPID 14059 8959 0.02 28
MOD_RES 268774 75331 0.47 7
MOTIF 49441 31809 0.09 21
MUTAGEN 108262 21627 0.19 16
NON_CONS 2641 829 <0.01 35
NON_STD 360 285 <0.01 36
NON_TER 12570 9667 0.02 30
PEPTIDE 12886 8910 0.02 29
PROPEP 15633 13364 0.03 27
REGION 329961 151799 0.57 6
REPEAT 110205 15302 0.19 15
SIGNAL 45273 45272 0.08 22
SITE 69435 37261 0.12 19
STRAND 378775 29914 0.66 4
TOPO_DOM 154982 31064 0.27 11
TRANSIT 9674 9551 0.02 32
TRANSMEM 385580 80748 0.67 3
TURN 90810 26005 0.16 18
UNSURE 5773 909 0.01 33
VAR_SEQ 53476 22769 0.09 20
VARIANT 106543 17640 0.19 17
ZN_FING 30967 13240 0.05 23
Total number of feature keys: 36
Total Number of Average
Line type / subtype number entries per entry Rank Category
------------------------------------ -------- --------- --------- ---- -------------------------------------------
Cross-references (DR) 21823477 37.98
ABCD 3196 3196 0.01 123 Protocols and materials databases
Agora 18462 18413 0.03 86 Miscellaneous databases
AGR 69344 68617 0.12 43 Organism-specific databases
Allergome 2047 1316 <0.01 133 Protein family/group databases
AlphaFoldDB 549408 549408 0.96 10 3D structure databases
Antibodypedia 32354 32245 0.06 62 Protocols and materials databases
AntiFam 22 22 <0.01 169 Family and domain databases
ArachnoServer 1148 1138 <0.01 141 Organism-specific databases
Araport 16438 16342 0.03 92 Organism-specific databases
Bgee 61985 61984 0.11 45 Gene expression databases
BindingDB 6929 6929 0.01 109 Chemistry databases
BioCyc 48261 44210 0.08 55 Enzyme and pathway databases
BioGRID 62481 60445 0.11 44 Protein-protein interaction databases
BioGRID-ORCS 45145 44557 0.08 56 Miscellaneous databases
BioMuta 20286 20259 0.04 74 Genetic variation databases
BMRB 6914 6914 0.01 110 3D structure databases
BRENDA 20504 18683 0.04 71 Enzyme and pathway databases
CarbonylDB 1159 1159 <0.01 140 PTM databases
CARD 321 319 <0.01 158 Protein family/group databases
CAZy 9723 8755 0.02 100 Protein family/group databases
CCDS 49766 34842 0.09 52 Sequence databases
CD-CODE 10734 8219 0.02 98 Miscellaneous databases
CDD 393904 309774 0.69 17 Family and domain databases
CGD 2160 2143 <0.01 131 Organism-specific databases
ChEMBL 9290 9110 0.02 101 Chemistry databases
ChiTaRS 15285 15251 0.03 95 Miscellaneous databases
CIViC 569 568 <0.01 150 Organism-specific databases
ClinPGx 18028 18009 0.03 88 Organism-specific databases
CollecTF 138 138 <0.01 161 Gene expression databases
ComplexPortal 19302 9733 0.03 82 Protein-protein interaction databases
ConoServer 967 879 <0.01 144 Organism-specific databases
CORUM 8090 8090 0.01 105 Protein-protein interaction databases
CPTAC 3472 1929 0.01 119 Proteomic databases
CPTC 410 410 <0.01 153 Protocols and materials databases
CTD 77698 77047 0.14 41 Organism-specific databases
DEPOD 254 254 <0.01 160 PTM databases
dictyBase 4228 4114 0.01 116 Organism-specific databases
DIP 17580 17539 0.03 90 Protein-protein interaction databases
DisGeNET 17613 17415 0.03 89 Organism-specific databases
DisProt 2825 2800 <0.01 126 Family and domain databases
DMDM 16164 16163 0.03 94 Genetic variation databases
DNASU 48556 48477 0.08 54 Protocols and materials databases
DrugBank 35601 4939 0.06 61 Chemistry databases
DrugCentral 2982 2982 0.01 125 Chemistry databases
EchoBASE 4158 4158 0.01 117 Organism-specific databases
eggNOG 340513 334622 0.59 20 Phylogenomic databases
ELM 1815 1815 <0.01 134 Protein-protein interaction databases
EMBL 1011147 561609 1.76 3 Sequence databases
EMDB 127943 11682 0.22 32 3D structure databases
Ensembl 122986 51864 0.21 34 Genome annotation databases
EnsemblBacteria 55578 55400 0.10 48 Genome annotation databases
EnsemblFungi 19513 19196 0.03 79 Genome annotation databases
EnsemblMetazoa 19685 12907 0.03 78 Genome annotation databases
EnsemblPlants 26391 6359 0.05 66 Genome annotation databases
EnsemblProtists 1740 1593 <0.01 135 Genome annotation databases
ESTHER 3035 3032 0.01 124 Protein family/group databases
euHCVdb 55 44 <0.01 166 Organism-specific databases
EvolutionaryTrace 22703 22703 0.04 69 Miscellaneous databases
ExpressionAtlas 51312 51312 0.09 50 Gene expression databases
FlyBase 3997 3888 0.01 118 Organism-specific databases
FunCoup 143520 143520 0.25 30 Protein-protein interaction databases
FunFam 558753 327653 0.97 9 Family and domain databases
Gene3D 800342 478318 1.39 6 Family and domain databases
GeneCards 20377 20247 0.04 73 Organism-specific databases
GeneID 315366 288167 0.55 23 Genome annotation databases
GeneReviews 1623 1619 <0.01 136 Organism-specific databases
GeneTree 48938 48921 0.09 53 Phylogenomic databases
GeneWiki 10351 10269 0.02 99 Miscellaneous databases
GenomeRNAi 22327 22326 0.04 70 Miscellaneous databases
GlyConnect 2372 2215 <0.01 128 PTM databases
GlyCosmos 28908 28908 0.05 64 PTM databases
GlyGen 39091 39091 0.07 59 PTM databases
GO 3358100 553982 5.84 1 Ontologies
Gramene 51809 22301 0.09 49 Genome annotation databases
GuidetoPHARMACOLOGY 2299 2299 <0.01 130 Chemistry databases
HAMAP 331036 328099 0.58 22 Family and domain databases
HGNC 20382 20256 0.04 72 Organism-specific databases
HOGENOM 428629 428629 0.75 16 Phylogenomic databases
HPA 19354 19215 0.03 81 Organism-specific databases
IDEAL 1101 1101 <0.01 142 Family and domain databases
IMGT_GENE-DB 267 267 <0.01 159 Protein family/group databases
InParanoid 164709 164709 0.29 26 Phylogenomic databases
IntAct 57960 57960 0.10 46 Protein-protein interaction databases
InterPro 2597857 556937 4.52 2 Family and domain databases
iPTMnet 56791 56791 0.10 47 PTM databases
JaponicusDB 43 43 <0.01 167 Organism-specific databases
jPOST 29053 29053 0.05 63 Proteomic databases
KEGG 520948 484511 0.91 13 Genome annotation databases
LegioList 765 763 <0.01 147 Organism-specific databases
Leproma 672 669 <0.01 148 Organism-specific databases
MaizeGDB 529 525 <0.01 151 Organism-specific databases
MalaCards 7396 7383 0.01 107 Organism-specific databases
MANE-Select 18592 18480 0.03 85 Genome annotation databases
MassIVE 19140 19140 0.03 83 Proteomic databases
MEROPS 14264 13845 0.02 97 Protein family/group databases
MetOSite 3455 3455 0.01 120 PTM databases
MGI 17168 17125 0.03 91 Organism-specific databases
MIM 24144 16578 0.04 68 Organism-specific databases
MINT 24151 24151 0.04 67 Protein-protein interaction databases
MoonDB 348 348 <0.01 157 Protein family/group databases
MoonProt 368 368 <0.01 155 Protein family/group databases
NCBIfam 547439 347231 0.95 11 Family and domain databases
NIAGADS 76 76 <0.01 163 Organism-specific databases
OGP 373 373 <0.01 154 2D gel databases
OMA 120955 120955 0.21 35 Phylogenomic databases
OpenTargets 18614 18469 0.03 84 Organism-specific databases
Orphanet 8157 4441 0.01 104 Organism-specific databases
OrthoDB 270718 270718 0.47 24 Phylogenomic databases
PAN-GO 20210 20210 0.04 75 Phylogenomic databases
PANTHER 964680 505734 1.68 4 Family and domain databases
PathwayCommons 19433 19433 0.03 80 Enzyme and pathway databases
PATRIC 93326 93326 0.16 39 Genome annotation databases
PaxDb 154216 154216 0.27 27 Proteomic databases
PCDDB 134 134 <0.01 162 3D structure databases
PDB 361254 38379 0.63 19 3D structure databases
PDBsum 361254 38379 0.63 18 3D structure databases
PeptideAtlas 38922 38922 0.07 60 Proteomic databases
PeroxiBase 794 773 <0.01 146 Protein family/group databases
Pfam 871800 547272 1.52 5 Family and domain databases
Pharos 20192 20192 0.04 76 Miscellaneous databases
PHI-base 2470 1949 <0.01 127 Miscellaneous databases
PhosphoSitePlus 42259 42259 0.07 58 PTM databases
PhylomeDB 115823 115823 0.20 36 Phylogenomic databases
PIR 125284 114939 0.22 33 Sequence databases
PIRSF 111097 109925 0.19 37 Family and domain databases
PlantReactome 1436 824 <0.01 138 Enzyme and pathway databases
PomBase 5135 5131 0.01 114 Organism-specific databases
PRIDE 637 637 <0.01 149 Proteomic databases
PRINTS 151585 130114 0.26 28 Family and domain databases
PRO 100156 100156 0.17 38 Miscellaneous databases
ProMEX 489 489 <0.01 152 Proteomic databases
PROSITE 496275 313384 0.86 14 Family and domain databases
Proteomes 491388 450492 0.86 15 Miscellaneous databases
ProteomicsDB 72937 45501 0.13 42 Proteomic databases
PseudoCAP 2054 2054 <0.01 132 Organism-specific databases
Pumba 18203 18203 0.03 87 Proteomic databases
Reactome 148027 39443 0.26 29 Enzyme and pathway databases
REBASE 802 395 <0.01 145 Protein family/group databases
RefSeq 566363 442362 0.99 8 Sequence databases
REPRODUCTION-2DPAGE 1260 1039 <0.01 139 2D gel databases
RGD 8159 8158 0.01 103 Organism-specific databases
RNAct 43120 43120 0.08 57 Miscellaneous databases
SABIO-RK 5951 5951 0.01 112 Enzyme and pathway databases
SASBDB 1025 1025 <0.01 143 3D structure databases
SFLD 27770 9138 0.05 65 Family and domain databases
SGD 6753 6748 0.01 111 Organism-specific databases
SignaLink 19948 19948 0.03 77 Enzyme and pathway databases
SIGNOR 7769 7769 0.01 106 Enzyme and pathway databases
SMART 207127 149377 0.36 25 Family and domain databases
SMR 525917 525917 0.92 12 3D structure databases
STRENDA-DB 59 45 <0.01 164 Enzyme and pathway databases
STRING 337189 337189 0.59 21 Protein-protein interaction databases
SUPFAM 652113 461938 1.13 7 Family and domain databases
SwissLipids 1478 1394 <0.01 137 Chemistry databases
SwissPalm 14331 14331 0.02 96 PTM databases
TAIR 16432 16342 0.03 93 Organism-specific databases
TCDB 8821 8723 0.02 102 Protein family/group databases
TopDownProteomics 3236 2957 0.01 122 Proteomic databases
TubercuList 2359 2323 <0.01 129 Organism-specific databases
UCSC 51124 46623 0.09 51 Genome annotation databases
UniLectin 367 367 <0.01 156 Protein family/group databases
UniPathway 140553 126895 0.24 31 Enzyme and pathway databases
VEuPathDB 87783 80328 0.15 40 Organism-specific databases
VGNC 5155 5143 0.01 113 Organism-specific databases
WBParaSite 56 54 <0.01 165 Genome annotation databases
WormBase 6951 5108 0.01 108 Organism-specific databases
Xenbase 4758 4758 0.01 115 Organism-specific databases
YCharOS 36 36 <0.01 168 Protocols and materials databases
ZFIN 3298 3295 0.01 121 Organism-specific databases
Total number of cross-referenced databases: 169
6. AMINO ACID COMPOSITION
6.1 Composition in percent for the complete database
Ala (A) 8.25 Gln (Q) 3.93 Leu (L) 9.64 Ser (S) 6.66
Arg (R) 5.52 Glu (E) 6.71 Lys (K) 5.79 Thr (T) 5.36
Asn (N) 4.06 Gly (G) 7.07 Met (M) 2.41 Trp (W) 1.10
Asp (D) 5.46 His (H) 2.27 Phe (F) 3.86 Tyr (Y) 2.92
Cys (C) 1.38 Ile (I) 5.90 Pro (P) 4.74 Val (V) 6.85
Asx (B) 0.000 Glx (Z) 0.000 Xaa (X) 0.00
The average sequence length in UniProtKB/Swiss-Prot is 362 amino acids.
The shortest sequence is GWA_SEPOF (P83570): 2 amino acids.
The longest sequence is TITIN_MOUSE (A2ASS6): 35213 amino acids.
4. JOURNAL CITATIONS
Note: the following citation statistics reflect the number of distinct
journal citations.
Total number of journals cited in this release of UniProtKB/Swiss-Prot: 3256
4.1 Table of the frequency of journal citations
Journals cited 1x: 1018
2x: 432
3x: 233
4x: 154
5x: 128
6x: 102
7x: 64
8x: 70
9x: 63
10x: 36
11- 20x: 257
21- 50x: 283
51-100x: 144
>100x: 272
4.2 List of the most cited journals in UniProtKB/Swiss-Prot
Nb Citations Journal name
-- --------- -------------------------------------------------------------
1 27952 Journal of Biological Chemistry
2 13174 Proceedings of the National Academy of Sciences of the U.S.A.
3 7396 Journal of Bacteriology
4 6197 Biochemical and Biophysical Research Communications
5 6014 Biochemistry
6 5511 Nucleic Acids Research
7 5415 Nature
8 5172 FEBS Letters
9 5115 The EMBO Journal
10 4911 Gene
11 4725 Journal of Molecular Biology
12 4667 Molecular and Cellular Biology
13 4121 Biochimica et Biophysica Acta
14 3985 Cell
15 3712 Journal of Virology
16 3538 European Journal of Biochemistry
17 3513 Science
18 3251 Biochemical Journal
19 3002 Molecular Microbiology
20 2844 Plant Physiology
21 2829 PLoS ONE
22 2549 Genomics
23 2501 The American Journal of Human Genetics
24 2432 Journal of Cell Biology
25 2217 The Plant Cell
26 2079 Human Molecular Genetics
27 2049 The Plant Journal
28 1994 Genes and Development
29 1976 Molecular Cell
30 1947 Virology
31 1929 Plant Molecular Biology
32 1875 Nature Genetics
33 1871 Molecular Biology of the Cell
34 1859 Development
35 1774 Journal of Immunology
36 1739 Nature Communications
37 1696 Human Mutation
38 1578 Oncogene
39 1519 Structure
40 1452 Journal of Biochemistry
41 1452 Genetics
42 1444 Molecular and General Genetics
43 1440 Journal of Cell Science
44 1327 Blood
45 1302 Infection and Immunity
46 1220 Microbiology
47 1212 Developmental Biology
48 1200 Journal of General Virology
49 1179 Current Biology
50 1172 Archives of Biochemistry and Biophysics
51 1099 Scientific Reports
52 1078 Journal of Neuroscience
53 1069 Applied and Environmental Microbiology
54 1013 Acta Crystallographica, Section D
55 973 PLoS Genetics
56 963 FEMS Microbiology Letters
57 940 Cancer Research
58 912 American Journal of Physiology
59 905 Toxicon
60 903 Protein Science
61 892 Journal of Clinical Investigation
62 860 Yeast
63 856 Neuron
64 807 The Journal of Experimental Medicine
65 783 PLoS Pathogens
66 779 Human Genetics
67 776 Plant and Cell Physiology
68 775 Nature Structural and Molecular Biology
69 745 Journal of Medical Genetics
70 740 The FEBS Journal
71 715 Proteins
72 684 Nature Cell Biology
73 682 Mechanisms of Development
74 661 Bioscience, Biotechnology, and Biochemistry
75 656 Nature Structural Biology
76 649 Antimicrobial Agents and Chemotherapy
77 618 Cell Reports
78 617 Developmental Cell
79 611 Current Genetics
80 584 Journal of Neurochemistry
81 564 Journal of the American Chemical Society
82 561 Molecular Endocrinology
83 558 The Journal of Clinical Endocrinology and Metabolism
84 555 Endocrinology
85 549 Molecular and Biochemical Parasitology
86 543
87 520 Eukaryotic Cell
88 505 Experimental Cell Research
89 504 EMBO Reports
90 499 RNA
91 495 Mammalian Genome
92 490 American Journal of Medical Genetics. Part A
93 486 The FASEB Journal
94 483 Peptides
95 477 Journal of Experimental Botany
96 462 Planta
97 449 Molecular Pharmacology
98 445 Acta Crystallographica, Section F
99 444 European Journal of Human Genetics
100 437 Clinical Genetics
101 435 Immunogenetics
102 432 Molecular Plant-Microbe Interactions
103 426 Immunity
104 426 Molecular Biology and Evolution
105 422 Journal of Investigative Dermatology
106 407 Journal of Molecular Evolution
107 402 Neurology
108 400 Biochimie
109 398 DNA and Cell Biology
110 387 Biology of Reproduction
111 381 Comparative Biochemistry and Physiology
112 381 DNA Sequence
113 376 PLoS Biology
114 372 Genes to Cells
115 368 Applied Microbiology and Biotechnology
116 366 Nature Immunology
117 366 Virus Research
118 359 Journal of Lipid Research
119 356 Journal of Medicinal Chemistry
120 352 BMC Genomics
121 350 Developmental Dynamics
122 346 The New England Journal of Medicine
123 343 Brain Research. Molecular Brain Research
124 341 Annals of Neurology
125 329 European Journal of Immunology
126 321 Journal of Human Genetics
127 321 Genome Research
128 315 Nature Chemical Biology
129 315 Investigative Ophthalmology and Visual Science
130 301 Brain
131 300 Glycobiology
132 299 Biological Chemistry Hoppe-Seyler
133 286 Fungal Genetics and Biology
134 285 Journal of General Microbiology
135 285 Archives of Microbiology
136 281 Cytogenetics and Cell Genetics
137 271 Traffic
138 271 Protein Expression and Purification
139 270 Cell Research
140 269 Molecular Genetics and Metabolism
141 266 Nature Medicine
142 263 Molecular Immunology
143 263 Phytochemistry
144 258 Journal of Cellular Biochemistry
145 254 Cell Cycle
146 246 Circulation Research
147 240 Insect Biochemistry and Molecular Biology
148 240 Diabetes
149 238 New Phytologist
150 237 Chemistry and Biology
5. STATISTICS FOR SOME LINE TYPES
The following table summarizes the total number of some UniProtKB/Swiss-Prot lines,
as well as the number of entries with at least one such line, and the
frequency of the lines.
Total Number of Average
Line type / subtype number entries per entry
------------------------------------ -------- --------- ---------
References (RL) 1338719 2.33
Journal 1167917 480978 2.03 1
Submitted to EMBL/GenBank/DDBJ 159128 143141 0.28 2
Submitted to other databases 7944 7239 0.01 3
Book citation 1877 1854 <0.01 4
Plant Gene Register 613 600 <0.01 5
Unpublished observations 543 539 <0.01 6
Thesis 477 474 <0.01 7
Patent 214 207 <0.01 8
Worm Breeder's Gazette 6 6 <0.01 9
Total number of distinct authors cited in UniProtKB/Swiss-Prot: 489697
Total Number of Average
Line type / subtype number entries per entry Rank
------------------------------------ -------- --------- --------- ----
Comments (CC) 2802336 4.88
ACTIVITY REGULATION 19539 19400 0.03 17
ALLERGEN 961 961 <0.01 26
ALTERNATIVE PRODUCTS 26020 26020 0.05 14
BIOPHYSICOCHEMICAL PROPERTIES 12354 12292 0.02 20
BIOTECHNOLOGY 2342 2281 <0.01 24
CATALYTIC ACTIVITY 357782 260363 0.62 4
CAUTION 14627 14324 0.03 19
COFACTOR 135473 123006 0.24 7
DEVELOPMENTAL STAGE 14912 14783 0.03 18
DISEASE 8698 5835 0.02 21
DISRUPTION PHENOTYPE 23450 23377 0.04 16
DOMAIN 62834 53157 0.11 9
FUNCTION 500305 473996 0.87 2
INDUCTION 27070 26950 0.05 13
INTERACTION 25638 25638 0.04 15
MASS SPECTROMETRY 7707 5971 0.01 22
MISCELLANEOUS 46679 41012 0.08 11
PATHWAY 145085 131041 0.25 6
PHARMACEUTICAL 169 162 <0.01 29
POLYMORPHISM 1515 1392 <0.01 25
PTM 67522 47537 0.12 8
RNA EDITING 646 646 <0.01 28
SEQUENCE CAUTION 45415 45345 0.08 12
SIMILARITY 522913 518545 0.91 1
SUBCELLULAR LOCATION 370136 361146 0.64 3
SUBUNIT 304021 298021 0.53 5
TISSUE SPECIFICITY 52275 51527 0.09 10
TOXIC DOSE 893 714 <0.01 27
WEB RESOURCE 5355 4823 0.01 23
Total number of comment topics: 29
Total Number of Average
Line type / subtype number entries per entry Rank
------------------------------------ -------- --------- --------- ----
Features (FT) 5659423 9.85
ACT_SITE 179698 107127 0.31 10
BINDING 1287551 222598 2.24 1
CARBOHYD 126559 32187 0.22 14
CHAIN 583134 566838 1.01 2
COILED 22877 15826 0.04 25
COMPBIAS 267854 93109 0.47 8
CONFLICT 140100 48791 0.24 13
CROSSLNK 25906 9283 0.05 24
DISULFID 141462 37439 0.25 12
DNA_BIND 12308 11027 0.02 31
DOMAIN 220216 134721 0.38 9
HELIX 374823 31767 0.65 5
INIT_MET 17661 17607 0.03 26
INTRAMEM 3194 1527 0.01 34
LIPID 14059 8959 0.02 28
MOD_RES 268774 75331 0.47 7
MOTIF 49441 31809 0.09 21
MUTAGEN 108262 21627 0.19 16
NON_CONS 2641 829 <0.01 35
NON_STD 360 285 <0.01 36
NON_TER 12570 9667 0.02 30
PEPTIDE 12886 8910 0.02 29
PROPEP 15633 13364 0.03 27
REGION 329961 151799 0.57 6
REPEAT 110205 15302 0.19 15
SIGNAL 45273 45272 0.08 22
SITE 69435 37261 0.12 19
STRAND 378775 29914 0.66 4
TOPO_DOM 154982 31064 0.27 11
TRANSIT 9674 9551 0.02 32
TRANSMEM 385580 80748 0.67 3
TURN 90810 26005 0.16 18
UNSURE 5773 909 0.01 33
VAR_SEQ 53476 22769 0.09 20
VARIANT 106543 17640 0.19 17
ZN_FING 30967 13240 0.05 23
Total number of feature keys: 36
Total Number of Average
Line type / subtype number entries per entry Rank Category
------------------------------------ -------- --------- --------- ---- -------------------------------------------
Cross-references (DR) 21823477 37.98
ABCD 3196 3196 0.01 123 Protocols and materials databases
Agora 18462 18413 0.03 86 Miscellaneous databases
AGR 69344 68617 0.12 43 Organism-specific databases
Allergome 2047 1316 <0.01 133 Protein family/group databases
AlphaFoldDB 549408 549408 0.96 10 3D structure databases
Antibodypedia 32354 32245 0.06 62 Protocols and materials databases
AntiFam 22 22 <0.01 169 Family and domain databases
ArachnoServer 1148 1138 <0.01 141 Organism-specific databases
Araport 16438 16342 0.03 92 Organism-specific databases
Bgee 61985 61984 0.11 45 Gene expression databases
BindingDB 6929 6929 0.01 109 Chemistry databases
BioCyc 48261 44210 0.08 55 Enzyme and pathway databases
BioGRID 62481 60445 0.11 44 Protein-protein interaction databases
BioGRID-ORCS 45145 44557 0.08 56 Miscellaneous databases
BioMuta 20286 20259 0.04 74 Genetic variation databases
BMRB 6914 6914 0.01 110 3D structure databases
BRENDA 20504 18683 0.04 71 Enzyme and pathway databases
CarbonylDB 1159 1159 <0.01 140 PTM databases
CARD 321 319 <0.01 158 Protein family/group databases
CAZy 9723 8755 0.02 100 Protein family/group databases
CCDS 49766 34842 0.09 52 Sequence databases
CD-CODE 10734 8219 0.02 98 Miscellaneous databases
CDD 393904 309774 0.69 17 Family and domain databases
CGD 2160 2143 <0.01 131 Organism-specific databases
ChEMBL 9290 9110 0.02 101 Chemistry databases
ChiTaRS 15285 15251 0.03 95 Miscellaneous databases
CIViC 569 568 <0.01 150 Organism-specific databases
ClinPGx 18028 18009 0.03 88 Organism-specific databases
CollecTF 138 138 <0.01 161 Gene expression databases
ComplexPortal 19302 9733 0.03 82 Protein-protein interaction databases
ConoServer 967 879 <0.01 144 Organism-specific databases
CORUM 8090 8090 0.01 105 Protein-protein interaction databases
CPTAC 3472 1929 0.01 119 Proteomic databases
CPTC 410 410 <0.01 153 Protocols and materials databases
CTD 77698 77047 0.14 41 Organism-specific databases
DEPOD 254 254 <0.01 160 PTM databases
dictyBase 4228 4114 0.01 116 Organism-specific databases
DIP 17580 17539 0.03 90 Protein-protein interaction databases
DisGeNET 17613 17415 0.03 89 Organism-specific databases
DisProt 2825 2800 <0.01 126 Family and domain databases
DMDM 16164 16163 0.03 94 Genetic variation databases
DNASU 48556 48477 0.08 54 Protocols and materials databases
DrugBank 35601 4939 0.06 61 Chemistry databases
DrugCentral 2982 2982 0.01 125 Chemistry databases
EchoBASE 4158 4158 0.01 117 Organism-specific databases
eggNOG 340513 334622 0.59 20 Phylogenomic databases
ELM 1815 1815 <0.01 134 Protein-protein interaction databases
EMBL 1011147 561609 1.76 3 Sequence databases
EMDB 127943 11682 0.22 32 3D structure databases
Ensembl 122986 51864 0.21 34 Genome annotation databases
EnsemblBacteria 55578 55400 0.10 48 Genome annotation databases
EnsemblFungi 19513 19196 0.03 79 Genome annotation databases
EnsemblMetazoa 19685 12907 0.03 78 Genome annotation databases
EnsemblPlants 26391 6359 0.05 66 Genome annotation databases
EnsemblProtists 1740 1593 <0.01 135 Genome annotation databases
ESTHER 3035 3032 0.01 124 Protein family/group databases
euHCVdb 55 44 <0.01 166 Organism-specific databases
EvolutionaryTrace 22703 22703 0.04 69 Miscellaneous databases
ExpressionAtlas 51312 51312 0.09 50 Gene expression databases
FlyBase 3997 3888 0.01 118 Organism-specific databases
FunCoup 143520 143520 0.25 30 Protein-protein interaction databases
FunFam 558753 327653 0.97 9 Family and domain databases
Gene3D 800342 478318 1.39 6 Family and domain databases
GeneCards 20377 20247 0.04 73 Organism-specific databases
GeneID 315366 288167 0.55 23 Genome annotation databases
GeneReviews 1623 1619 <0.01 136 Organism-specific databases
GeneTree 48938 48921 0.09 53 Phylogenomic databases
GeneWiki 10351 10269 0.02 99 Miscellaneous databases
GenomeRNAi 22327 22326 0.04 70 Miscellaneous databases
GlyConnect 2372 2215 <0.01 128 PTM databases
GlyCosmos 28908 28908 0.05 64 PTM databases
GlyGen 39091 39091 0.07 59 PTM databases
GO 3358100 553982 5.84 1 Ontologies
Gramene 51809 22301 0.09 49 Genome annotation databases
GuidetoPHARMACOLOGY 2299 2299 <0.01 130 Chemistry databases
HAMAP 331036 328099 0.58 22 Family and domain databases
HGNC 20382 20256 0.04 72 Organism-specific databases
HOGENOM 428629 428629 0.75 16 Phylogenomic databases
HPA 19354 19215 0.03 81 Organism-specific databases
IDEAL 1101 1101 <0.01 142 Family and domain databases
IMGT_GENE-DB 267 267 <0.01 159 Protein family/group databases
InParanoid 164709 164709 0.29 26 Phylogenomic databases
IntAct 57960 57960 0.10 46 Protein-protein interaction databases
InterPro 2597857 556937 4.52 2 Family and domain databases
iPTMnet 56791 56791 0.10 47 PTM databases
JaponicusDB 43 43 <0.01 167 Organism-specific databases
jPOST 29053 29053 0.05 63 Proteomic databases
KEGG 520948 484511 0.91 13 Genome annotation databases
LegioList 765 763 <0.01 147 Organism-specific databases
Leproma 672 669 <0.01 148 Organism-specific databases
MaizeGDB 529 525 <0.01 151 Organism-specific databases
MalaCards 7396 7383 0.01 107 Organism-specific databases
MANE-Select 18592 18480 0.03 85 Genome annotation databases
MassIVE 19140 19140 0.03 83 Proteomic databases
MEROPS 14264 13845 0.02 97 Protein family/group databases
MetOSite 3455 3455 0.01 120 PTM databases
MGI 17168 17125 0.03 91 Organism-specific databases
MIM 24144 16578 0.04 68 Organism-specific databases
MINT 24151 24151 0.04 67 Protein-protein interaction databases
MoonDB 348 348 <0.01 157 Protein family/group databases
MoonProt 368 368 <0.01 155 Protein family/group databases
NCBIfam 547439 347231 0.95 11 Family and domain databases
NIAGADS 76 76 <0.01 163 Organism-specific databases
OGP 373 373 <0.01 154 2D gel databases
OMA 120955 120955 0.21 35 Phylogenomic databases
OpenTargets 18614 18469 0.03 84 Organism-specific databases
Orphanet 8157 4441 0.01 104 Organism-specific databases
OrthoDB 270718 270718 0.47 24 Phylogenomic databases
PAN-GO 20210 20210 0.04 75 Phylogenomic databases
PANTHER 964680 505734 1.68 4 Family and domain databases
PathwayCommons 19433 19433 0.03 80 Enzyme and pathway databases
PATRIC 93326 93326 0.16 39 Genome annotation databases
PaxDb 154216 154216 0.27 27 Proteomic databases
PCDDB 134 134 <0.01 162 3D structure databases
PDB 361254 38379 0.63 19 3D structure databases
PDBsum 361254 38379 0.63 18 3D structure databases
PeptideAtlas 38922 38922 0.07 60 Proteomic databases
PeroxiBase 794 773 <0.01 146 Protein family/group databases
Pfam 871800 547272 1.52 5 Family and domain databases
Pharos 20192 20192 0.04 76 Miscellaneous databases
PHI-base 2470 1949 <0.01 127 Miscellaneous databases
PhosphoSitePlus 42259 42259 0.07 58 PTM databases
PhylomeDB 115823 115823 0.20 36 Phylogenomic databases
PIR 125284 114939 0.22 33 Sequence databases
PIRSF 111097 109925 0.19 37 Family and domain databases
PlantReactome 1436 824 <0.01 138 Enzyme and pathway databases
PomBase 5135 5131 0.01 114 Organism-specific databases
PRIDE 637 637 <0.01 149 Proteomic databases
PRINTS 151585 130114 0.26 28 Family and domain databases
PRO 100156 100156 0.17 38 Miscellaneous databases
ProMEX 489 489 <0.01 152 Proteomic databases
PROSITE 496275 313384 0.86 14 Family and domain databases
Proteomes 491388 450492 0.86 15 Miscellaneous databases
ProteomicsDB 72937 45501 0.13 42 Proteomic databases
PseudoCAP 2054 2054 <0.01 132 Organism-specific databases
Pumba 18203 18203 0.03 87 Proteomic databases
Reactome 148027 39443 0.26 29 Enzyme and pathway databases
REBASE 802 395 <0.01 145 Protein family/group databases
RefSeq 566363 442362 0.99 8 Sequence databases
REPRODUCTION-2DPAGE 1260 1039 <0.01 139 2D gel databases
RGD 8159 8158 0.01 103 Organism-specific databases
RNAct 43120 43120 0.08 57 Miscellaneous databases
SABIO-RK 5951 5951 0.01 112 Enzyme and pathway databases
SASBDB 1025 1025 <0.01 143 3D structure databases
SFLD 27770 9138 0.05 65 Family and domain databases
SGD 6753 6748 0.01 111 Organism-specific databases
SignaLink 19948 19948 0.03 77 Enzyme and pathway databases
SIGNOR 7769 7769 0.01 106 Enzyme and pathway databases
SMART 207127 149377 0.36 25 Family and domain databases
SMR 525917 525917 0.92 12 3D structure databases
STRENDA-DB 59 45 <0.01 164 Enzyme and pathway databases
STRING 337189 337189 0.59 21 Protein-protein interaction databases
SUPFAM 652113 461938 1.13 7 Family and domain databases
SwissLipids 1478 1394 <0.01 137 Chemistry databases
SwissPalm 14331 14331 0.02 96 PTM databases
TAIR 16432 16342 0.03 93 Organism-specific databases
TCDB 8821 8723 0.02 102 Protein family/group databases
TopDownProteomics 3236 2957 0.01 122 Proteomic databases
TubercuList 2359 2323 <0.01 129 Organism-specific databases
UCSC 51124 46623 0.09 51 Genome annotation databases
UniLectin 367 367 <0.01 156 Protein family/group databases
UniPathway 140553 126895 0.24 31 Enzyme and pathway databases
VEuPathDB 87783 80328 0.15 40 Organism-specific databases
VGNC 5155 5143 0.01 113 Organism-specific databases
WBParaSite 56 54 <0.01 165 Genome annotation databases
WormBase 6951 5108 0.01 108 Organism-specific databases
Xenbase 4758 4758 0.01 115 Organism-specific databases
YCharOS 36 36 <0.01 168 Protocols and materials databases
ZFIN 3298 3295 0.01 121 Organism-specific databases
Total number of cross-referenced databases: 169
6. AMINO ACID COMPOSITION
6.1 Composition in percent for the complete database
Ala (A) 8.25 Gln (Q) 3.93 Leu (L) 9.64 Ser (S) 6.66
Arg (R) 5.52 Glu (E) 6.71 Lys (K) 5.79 Thr (T) 5.36
Asn (N) 4.06 Gly (G) 7.07 Met (M) 2.41 Trp (W) 1.10
Asp (D) 5.46 His (H) 2.27 Phe (F) 3.86 Tyr (Y) 2.92
Cys (C) 1.38 Ile (I) 5.90 Pro (P) 4.74 Val (V) 6.85
Asx (B) 0.000 Glx (Z) 0.000 Xaa (X) 0.00
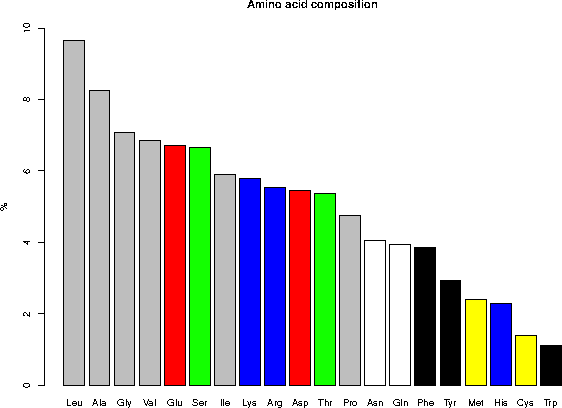 Legend: gray = aliphatic, red = acidic, green = small hydroxy,
blue = basic, black = aromatic, white = amide, yellow = sulfur
6.2 Classification of the amino acids by their frequency
Leu, Ala, Gly, Val, Glu, Ser, Ile, Lys, Arg, Asp, Thr, Pro, Asn, Gln,
Phe, Tyr, Met, His, Cys, Trp
7. MISCELLANEOUS STATISTICS
4473 entries are encoded on a mitochondrion, and 4063 are encoded on a plasmid.
12200 entries are encoded on a plastid,
of which 23 are encoded on apicoplasts,
11633 on chloroplasts,
51 on organellar chromatophores,
145 on cyanelles,
149 on non-photosynthetic plastids and
199 on unspecified types of plastid.
Number of entries with at least one sequence correction: 81583
Legend: gray = aliphatic, red = acidic, green = small hydroxy,
blue = basic, black = aromatic, white = amide, yellow = sulfur
6.2 Classification of the amino acids by their frequency
Leu, Ala, Gly, Val, Glu, Ser, Ile, Lys, Arg, Asp, Thr, Pro, Asn, Gln,
Phe, Tyr, Met, His, Cys, Trp
7. MISCELLANEOUS STATISTICS
4473 entries are encoded on a mitochondrion, and 4063 are encoded on a plasmid.
12200 entries are encoded on a plastid,
of which 23 are encoded on apicoplasts,
11633 on chloroplasts,
51 on organellar chromatophores,
145 on cyanelles,
149 on non-photosynthetic plastids and
199 on unspecified types of plastid.
Number of entries with at least one sequence correction: 81583
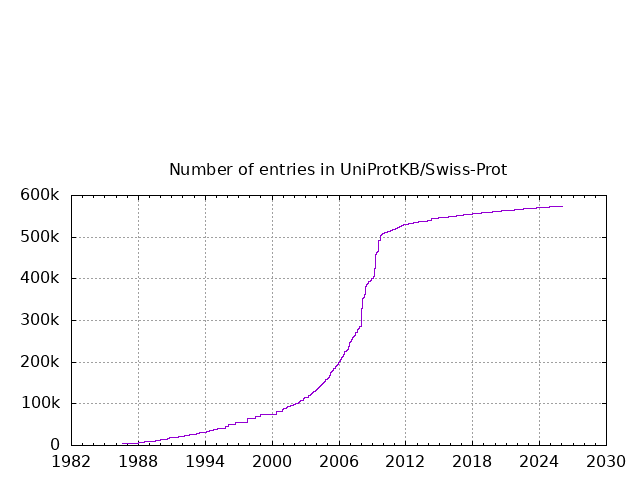 2. TAXONOMIC ORIGIN
Total number of species represented in this release of UniProtKB/Swiss-Prot: 14846
The first twenty species represent 123389 sequences: 21.5 % of the total
number of entries.
2.1 Table of the frequency of occurrence of species
Species represented 1x: 6040
2x: 2144
3x: 1167
4x: 793
5x: 549
6x: 448
7x: 332
8x: 295
9x: 238
10x: 162
11- 20x: 860
21- 50x: 521
51-100x: 235
>100x: 1062
2.2 Table of the most represented species
------ --------- --------------------------------------------
Number Frequency Species
------ --------- --------------------------------------------
1 20431 Homo sapiens (Human)
2 17252 Mus musculus (Mouse)
3 16418 Arabidopsis thaliana (Mouse-ear cress)
4 8226 Rattus norvegicus (Rat)
5 6733 Saccharomyces cerevisiae (strain ATCC 204508 / S288c) (Baker's yeast)
6 6052 Bos taurus (Bovine)
7 5129 Schizosaccharomyces pombe (strain 972 / ATCC 24843) (Fission yeast)
8 4531 Escherichia coli (strain K12)
9 4499 Caenorhabditis elegans
10 4197 Oryza sativa subsp. japonica (Rice)
11 4191 Bacillus subtilis (strain 168)
12 4163 Dictyostelium discoideum (Social amoeba)
13 3868 Drosophila melanogaster (Fruit fly)
14 3514 Xenopus laevis (African clawed frog)
15 3369 Danio rerio (Zebrafish) (Brachydanio rerio)
16 2338 Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv)
17 2314 Gallus gallus (Chicken)
18 2218 Pongo abelii (Sumatran orangutan) (Pongo pygmaeus abelii)
19 2047 Escherichia coli O157:H7
20 1899 Mycobacterium tuberculosis (strain CDC 1551 / Oshkosh)
21 1831 Salmonella typhimurium (strain LT2 / SGSC1412 / ATCC 700720)
22 1787 Methanocaldococcus jannaschii
23 1713 Xenopus tropicalis (Western clawed frog) (Silurana tropicalis)
24 1703 Haemophilus influenzae (strain ATCC 51907 / DSM 11121 / KW20 / Rd)
25 1702 Escherichia coli O6:H1 (strain CFT073 / ATCC 700928 / UPEC)
26 1696 Shigella flexneri
27 1479 Pseudomonas aeruginosa
28 1462 Sus scrofa (Pig)
29 1349 Salmonella typhi
30 1244 Mycobacterium bovis (strain ATCC BAA-935 / AF2122/97)
31 1197 Candida albicans (strain SC5314 / ATCC MYA-2876) (Yeast)
32 1176 Macaca fascicularis (Crab-eating macaque) (Cynomolgus monkey)
33 1108 Synechocystis sp. (strain ATCC 27184 / PCC 6803 / Kazusa)
34 1038 Archaeoglobus fulgidus
35 1030 Yersinia pestis
36 1028 Emericella nidulans
37 1000 Vibrio cholerae serotype O1 (strain ATCC 39315 / El Tor Inaba N16961)
38 979 Oryctolagus cuniculus (Rabbit)
39 970 Neurospora crassa
40 959 Aspergillus fumigatus (strain ATCC MYA-4609 / CBS 101355 / FGSC A1100 / Af293)
41 942 Staphylococcus aureus (strain Mu50 / ATCC 700699)
42 930 Salmonella paratyphi A (strain ATCC 9150 / SARB42)
43 929 Staphylococcus aureus (strain N315)
44 928 Eremothecium gossypii
45 920 Kluyveromyces lactis
46 909 Acanthamoeba polyphaga mimivirus (APMV)
47 905 Staphylococcus aureus (strain COL)
48 896 Staphylococcus aureus (strain MW2)
49 894 Escherichia coli O6:K15:H31 (strain 536 / UPEC)
50 892 Rhizobium meliloti (strain 1021) (Ensifer meliloti) (Sinorhizobium meliloti)
51 890 Candida glabrata
52 890 Staphylococcus aureus (strain MSSA476)
53 888 Staphylococcus aureus (strain MRSA252)
54 882 Salmonella choleraesuis (strain SC-B67)
55 879 Shigella sonnei (strain Ss046)
56 877 Oryza sativa subsp. indica (Rice)
57 863 Yersinia pseudotuberculosis serotype I (strain IP32953)
58 857 Canis lupus familiaris (Dog) (Canis familiaris)
59 850 Zea mays (Maize)
60 847 Escherichia coli O9:H4 (strain HS)
61 838 Escherichia coli O139:H28 (strain E24377A / ETEC)
62 829 Shigella boydii serotype 4 (strain Sb227)
63 825 Escherichia coli (strain UTI89 / UPEC)
64 822 Escherichia coli
65 822 Shigella dysenteriae serotype 1 (strain Sd197)
66 819 Streptomyces coelicolor (strain ATCC BAA-471 / A3(2) / M145)
67 813 Staphylococcus aureus (strain NCTC 8325 / PS 47)
68 804 Pectobacterium atrosepticum (strain SCRI 1043 / ATCC BAA-672)
69 796 Vibrio parahaemolyticus serotype O3:K6 (strain RIMD 2210633)
70 791 Escherichia coli (strain SMS-3-5 / SECEC)
71 788 Aquifex aeolicus (strain VF5)
72 779 Escherichia coli O127:H6 (strain E2348/69 / EPEC)
73 771 Escherichia coli (strain K12 / DH10B)
74 770 Pasteurella multocida (strain Pm70)
75 767 Escherichia coli O17:K52:H18 (strain UMN026 / ExPEC)
76 765 Escherichia coli (strain K12 / MC4100 / BW2952)
77 762 Escherichia coli (strain 55989 / EAEC)
78 761 Escherichia coli O8 (strain IAI1)
79 760 Shigella flexneri serotype 5b (strain 8401)
80 760 Staphylococcus epidermidis
81 760 Staphylococcus epidermidis (strain ATCC 12228 / FDA PCI 1200)
82 759 Escherichia coli O45:K1 (strain S88 / ExPEC)
83 758 Bacillus anthracis
84 756 Escherichia coli (strain SE11)
85 753 Escherichia coli O7:K1 (strain IAI39 / ExPEC)
86 749 Photorhabdus laumondii subsp. laumondii (strain DSM 15139 / CIP 105565 / TT01)
87 748 Escherichia coli O157:H7 (strain EC4115 / EHEC)
88 744 Halalkalibacterium halodurans
89 739 Yersinia enterocolitica serotype O:8 / biotype 1B (strain NCTC 13174 / 8081)
90 739 Escherichia coli
91 738 Pseudomonas putida
92 733 Vibrio vulnificus (strain CMCP6)
93 731 Escherichia coli O81 (strain ED1a)
94 722 Salmonella enteritidis PT4 (strain P125109)
95 719 Klebsiella pneumoniae subsp. pneumoniae (strain ATCC 700721 / MGH 78578)
96 718 Vibrio vulnificus (strain YJ016)
97 716 Salmonella paratyphi B (strain ATCC BAA-1250 / SPB7)
98 715 Enterobacter sp. (strain 638)
99 715 Escherichia coli O1:K1 / APEC
100 715 Yersinia pestis bv. Antiqua (strain Nepal516)
101 714 Salmonella paratyphi A (strain AKU_12601)
102 713 Yersinia pseudotuberculosis serotype O:1b (strain IP 31758)
103 713 Salmonella newport (strain SL254)
104 713 Salmonella agona (strain SL483)
105 712 Salmonella schwarzengrund (strain CVM19633)
106 711 Yersinia pestis bv. Antiqua (strain Antiqua)
107 710 Salmonella heidelberg (strain SL476)
108 708 Nostoc sp. (strain PCC 7120 / SAG 25.82 / UTEX 2576)
109 702 Salmonella dublin (strain CT_02021853)
110 699 Klebsiella variicola (strain 342) (Klebsiella pneumoniae)
111 698 Shigella boydii serotype 18 (strain CDC 3083-94 / BS512)
112 695 Escherichia fergusonii
113 692 Pan troglodytes (Chimpanzee)
114 686 Mycoplasma pneumoniae (strain ATCC 29342 / M129 / Subtype 1)
115 684 Salmonella gallinarum (strain 287/91 / NCTC 13346)
116 683 Pseudomonas syringae pv. tomato (strain ATCC BAA-871 / DC3000)
117 679 Agrobacterium fabrum (strain C58 / ATCC 33970) (Agrobacterium tumefaciens
118 679 Citrobacter koseri (strain ATCC BAA-895 / CDC 4225-83 / SGSC4696)
119 679 Staphylococcus aureus (strain USA300)
120 672 Serratia proteamaculans (strain 568)
121 672 Bacillus cereus
122 669 Mycobacterium leprae (strain TN)
123 667 Yarrowia lipolytica (strain CLIB 122 / E 150) (Yeast) (Candida lipolytica)
124 667 Bradyrhizobium diazoefficiens
125 667 Yersinia pestis (strain Pestoides F)
126 663 Shewanella oneidensis
127 658 Sinorhizobium fredii (strain NBRC 101917 / NGR234)
128 653 Debaryomyces hansenii
129 643 Staphylococcus aureus (strain bovine RF122 / ET3-1)
130 642 Yersinia pseudotuberculosis serotype O:3 (strain YPIII)
131 642 Salmonella arizonae (strain ATCC BAA-731 / CDC346-86 / RSK2980)
132 634 Yersinia pseudotuberculosis serotype IB (strain PB1/+)
133 623 Methanothermobacter thermautotrophicus
134 623 Cronobacter sakazakii (strain ATCC BAA-894) (Enterobacter sakazakii)
135 622 Treponema pallidum (strain Nichols)
136 622 Listeria monocytogenes serovar 1/2a (strain ATCC BAA-679 / EGD-e)
137 620 Pseudomonas aeruginosa (strain UCBPP-PA14)
138 615 Xanthomonas campestris pv. campestris
139 614 Mesorhizobium japonicum (Mesorhizobium loti
140 614 Staphylococcus haemolyticus (strain JCSC1435)
141 613 Helicobacter pylori (strain ATCC 700392 / 26695) (Campylobacter pylori)
142 611 Mycolicibacterium smegmatis (strain ATCC 700084 / mc(2)155)
143 605 Listeria innocua serovar 6a (strain ATCC BAA-680 / CLIP 11262)
144 604 Ralstonia nicotianae (strain ATCC BAA-1114 / GMI1000) (Ralstonia solanacearum)
145 602 Staphylococcus saprophyticus subsp. saprophyticus
146 602 Photobacterium profundum (strain SS9)
147 601 Salmonella paratyphi C (strain RKS4594)
148 600 Yersinia pestis bv. Antiqua (strain Angola)
149 595 Bacillus cereus (strain ATCC 10987 / NRS 248)
150 592 Neisseria meningitidis serogroup B (strain ATCC BAA-335 / MC58)
151 591 Pectobacterium carotovorum subsp. carotovorum (strain PC1)
152 588 Aspergillus oryzae (strain ATCC 42149 / RIB 40) (Yellow koji mold)
153 584 Rickettsia prowazekii (strain Madrid E)
154 582 Caenorhabditis briggsae
155 579 Brucella suis biovar 1 (strain 1330)
156 576 Brucella melitensis biotype 1
157 575 Caulobacter vibrioides (strain ATCC 19089 / CIP 103742 / CB 15)
158 573 Aliivibrio fischeri (strain ATCC 700601 / ES114) (Vibrio fischeri)
159 572 Buchnera aphidicola subsp. Acyrthosiphon pisum (strain APS)
160 569 Bacillus thuringiensis subsp. konkukian (strain 97-27)
161 568 Helicobacter pylori (strain J99 / ATCC 700824) (Campylobacter pylori J99)
162 568 Pseudomonas syringae pv. syringae (strain B728a)
163 567 Thermotoga maritima
164 566 Bacillus licheniformis
165 562 Bacillus cereus (strain ZK / E33L)
166 562 Buchnera aphidicola subsp. Schizaphis graminum (strain Sg)
167 561 Xanthomonas axonopodis pv. citri (strain 306)
168 559 Clostridium acetobutylicum
169 555 Pseudomonas fluorescens (strain Pf0-1)
170 554 Neisseria meningitidis serogroup A / serotype 4A (strain DSM 15465 / Z2491)
171 554 Pseudomonas fluorescens (strain ATCC BAA-477 / NRRL B-23932 / Pf-5)
172 553 Oceanobacillus iheyensis
173 547 Pseudomonas savastanoi pv. phaseolicola (Pseudomonas syringae pv. phaseolicola
174 543 Corynebacterium glutamicum
175 541 Lactococcus lactis subsp. lactis (strain IL1403) (Streptococcus lactis)
176 533 Bordetella bronchiseptica (strain ATCC BAA-588 / NCTC 13252 / RB50)
177 531 Erwinia tasmaniensis
178 530 Listeria monocytogenes serotype 4b (strain F2365)
179 529 Sodalis glossinidius (strain morsitans)
180 525 Staphylococcus aureus (strain Newman)
181 525 Deinococcus radiodurans
182 523 Vibrio cholerae serotype O1 (strain ATCC 39541 / Classical Ogawa 395 / O395)
183 522 Xylella fastidiosa (strain 9a5c)
184 519 Chromobacterium violaceum
185 519 Methanosarcina acetivorans (strain ATCC 35395 / DSM 2834 / JCM 12185 / C2A)
186 519 Streptococcus pneumoniae serotype 4 (strain ATCC BAA-334 / TIGR4)
187 516 Bordetella pertussis (strain Tohama I / ATCC BAA-589 / NCTC 13251)
188 515 Xylella fastidiosa (strain Temecula1 / ATCC 700964)
189 512 Pyrococcus furiosus (strain ATCC 43587 / DSM 3638 / JCM 8422 / Vc1)
190 512 Geobacillus kaustophilus (strain HTA426)
191 512 Haemophilus ducreyi (strain 35000HP / ATCC 700724)
192 512 Pseudomonas paraeruginosa (strain DSM 24068 / PA7) (Pseudomonas aeruginosa
193 511 Solanum lycopersicum (Tomato) (Lycopersicon esculentum)
194 511 Acinetobacter baylyi (strain ATCC 33305 / BD413 / ADP1)
195 511 Streptomyces avermitilis
196 509 Streptococcus pneumoniae (strain ATCC BAA-255 / R6)
197 508 Bordetella parapertussis (strain 12822 / ATCC BAA-587 / NCTC 13253)
198 507 Buchnera aphidicola subsp. Baizongia pistaciae (strain Bp)
199 506 Nicotiana tabacum (Common tobacco)
200 505 Saccharolobus solfataricus (strain ATCC 35092 / DSM 1617 / JCM 11322 / P2)
201 504 Pseudomonas entomophila (strain L48)
202 501 Methanosarcina mazei
203 501 Burkholderia pseudomallei (strain K96243)
204 499 Brucella abortus biovar 1 (strain 9-941)
205 499 Haemophilus influenzae (strain 86-028NP)
206 498 Thermosynechococcus vestitus (strain NIES-2133 / IAM M-273 / BP-1)
207 497 Xanthomonas campestris pv. campestris (strain 8004)
208 497 Synechococcus elongatus (strain ATCC 33912 / PCC 7942 / FACHB-805)
209 497 Pyrococcus horikoshii
210 497 Proteus mirabilis (strain HI4320)
211 496 Rickettsia conorii (strain ATCC VR-613 / Malish 7)
212 496 Shouchella clausii (strain KSM-K16) (Alkalihalobacillus clausii)
213 495 Halobacterium salinarum (strain ATCC 700922 / JCM 11081 / NRC-1)
214 493 Brucella abortus (strain 2308)
215 492 Bacillus velezensis (strain DSM 23117 / BGSC 10A6 / LMG 26770 / FZB42)
216 491 Vibrio campbellii (strain ATCC BAA-1116)
217 488 Shewanella sp. (strain MR-7)
218 486 Mannheimia succiniciproducens (strain KCTC 0769BP / MBEL55E)
219 485 Pseudomonas aeruginosa (strain LESB58)
220 485 Shewanella sp. (strain MR-4)
221 484 Staphylococcus aureus (strain Mu3 / ATCC 700698)
222 483 Mycoplasma genitalium (strain ATCC 33530 / DSM 19775 / NCTC 10195 / G37)
223 483 Lactiplantibacillus plantarum (strain ATCC BAA-793 / NCIMB 8826 / WCFS1)
224 480 Pseudomonas putida
225 478 Cupriavidus necator
226 478 Pyrococcus abyssi (strain GE5 / Orsay)
227 476 Enterococcus faecalis (strain ATCC 700802 / V583)
228 475 Burkholderia lata
229 475 Campylobacter jejuni subsp. jejuni serotype O:2
230 472 Rhodopseudomonas palustris (strain ATCC BAA-98 / CGA009)
231 471 Cereibacter sphaeroides
232 470 Clostridium perfringens (strain 13 / Type A)
233 468 Shewanella sp. (strain ANA-3)
234 468 Shewanella frigidimarina (strain NCIMB 400)
235 468 Pseudomonas putida (strain GB-1)
236 467 Aeromonas hydrophila subsp. hydrophila
237 466 Xanthomonas euvesicatoria pv. vesicatoria (strain 85-10)
238 465 Trichormus variabilis (strain ATCC 29413 / PCC 7937) (Anabaena variabilis)
239 463 Burkholderia mallei (strain ATCC 23344)
240 462 Cupriavidus pinatubonensis (strain JMP 134 / LMG 1197) (Cupriavidus necator
241 461 Ovis aries (Sheep)
242 460 Methylococcus capsulatus (strain ATCC 33009 / NCIMB 11132 / Bath)
243 457 Rickettsia felis (strain ATCC VR-1525 / URRWXCal2) (Rickettsia azadi)
244 455 Staphylococcus aureus (strain JH1)
245 455 Xanthomonas oryzae pv. oryzae (strain MAFF 311018)
246 455 Shewanella baltica (strain OS185)
247 453 Streptococcus mutans serotype c (strain ATCC 700610 / UA159)
248 453 Mycolicibacterium paratuberculosis (strain ATCC BAA-968 / K-10)
249 453 Pseudomonas putida (strain W619)
250 452 Caldanaerobacter subterraneus subsp. tengcongensis
2.3 Taxonomic distribution of the sequences
2. TAXONOMIC ORIGIN
Total number of species represented in this release of UniProtKB/Swiss-Prot: 14846
The first twenty species represent 123389 sequences: 21.5 % of the total
number of entries.
2.1 Table of the frequency of occurrence of species
Species represented 1x: 6040
2x: 2144
3x: 1167
4x: 793
5x: 549
6x: 448
7x: 332
8x: 295
9x: 238
10x: 162
11- 20x: 860
21- 50x: 521
51-100x: 235
>100x: 1062
2.2 Table of the most represented species
------ --------- --------------------------------------------
Number Frequency Species
------ --------- --------------------------------------------
1 20431 Homo sapiens (Human)
2 17252 Mus musculus (Mouse)
3 16418 Arabidopsis thaliana (Mouse-ear cress)
4 8226 Rattus norvegicus (Rat)
5 6733 Saccharomyces cerevisiae (strain ATCC 204508 / S288c) (Baker's yeast)
6 6052 Bos taurus (Bovine)
7 5129 Schizosaccharomyces pombe (strain 972 / ATCC 24843) (Fission yeast)
8 4531 Escherichia coli (strain K12)
9 4499 Caenorhabditis elegans
10 4197 Oryza sativa subsp. japonica (Rice)
11 4191 Bacillus subtilis (strain 168)
12 4163 Dictyostelium discoideum (Social amoeba)
13 3868 Drosophila melanogaster (Fruit fly)
14 3514 Xenopus laevis (African clawed frog)
15 3369 Danio rerio (Zebrafish) (Brachydanio rerio)
16 2338 Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv)
17 2314 Gallus gallus (Chicken)
18 2218 Pongo abelii (Sumatran orangutan) (Pongo pygmaeus abelii)
19 2047 Escherichia coli O157:H7
20 1899 Mycobacterium tuberculosis (strain CDC 1551 / Oshkosh)
21 1831 Salmonella typhimurium (strain LT2 / SGSC1412 / ATCC 700720)
22 1787 Methanocaldococcus jannaschii
23 1713 Xenopus tropicalis (Western clawed frog) (Silurana tropicalis)
24 1703 Haemophilus influenzae (strain ATCC 51907 / DSM 11121 / KW20 / Rd)
25 1702 Escherichia coli O6:H1 (strain CFT073 / ATCC 700928 / UPEC)
26 1696 Shigella flexneri
27 1479 Pseudomonas aeruginosa
28 1462 Sus scrofa (Pig)
29 1349 Salmonella typhi
30 1244 Mycobacterium bovis (strain ATCC BAA-935 / AF2122/97)
31 1197 Candida albicans (strain SC5314 / ATCC MYA-2876) (Yeast)
32 1176 Macaca fascicularis (Crab-eating macaque) (Cynomolgus monkey)
33 1108 Synechocystis sp. (strain ATCC 27184 / PCC 6803 / Kazusa)
34 1038 Archaeoglobus fulgidus
35 1030 Yersinia pestis
36 1028 Emericella nidulans
37 1000 Vibrio cholerae serotype O1 (strain ATCC 39315 / El Tor Inaba N16961)
38 979 Oryctolagus cuniculus (Rabbit)
39 970 Neurospora crassa
40 959 Aspergillus fumigatus (strain ATCC MYA-4609 / CBS 101355 / FGSC A1100 / Af293)
41 942 Staphylococcus aureus (strain Mu50 / ATCC 700699)
42 930 Salmonella paratyphi A (strain ATCC 9150 / SARB42)
43 929 Staphylococcus aureus (strain N315)
44 928 Eremothecium gossypii
45 920 Kluyveromyces lactis
46 909 Acanthamoeba polyphaga mimivirus (APMV)
47 905 Staphylococcus aureus (strain COL)
48 896 Staphylococcus aureus (strain MW2)
49 894 Escherichia coli O6:K15:H31 (strain 536 / UPEC)
50 892 Rhizobium meliloti (strain 1021) (Ensifer meliloti) (Sinorhizobium meliloti)
51 890 Candida glabrata
52 890 Staphylococcus aureus (strain MSSA476)
53 888 Staphylococcus aureus (strain MRSA252)
54 882 Salmonella choleraesuis (strain SC-B67)
55 879 Shigella sonnei (strain Ss046)
56 877 Oryza sativa subsp. indica (Rice)
57 863 Yersinia pseudotuberculosis serotype I (strain IP32953)
58 857 Canis lupus familiaris (Dog) (Canis familiaris)
59 850 Zea mays (Maize)
60 847 Escherichia coli O9:H4 (strain HS)
61 838 Escherichia coli O139:H28 (strain E24377A / ETEC)
62 829 Shigella boydii serotype 4 (strain Sb227)
63 825 Escherichia coli (strain UTI89 / UPEC)
64 822 Escherichia coli
65 822 Shigella dysenteriae serotype 1 (strain Sd197)
66 819 Streptomyces coelicolor (strain ATCC BAA-471 / A3(2) / M145)
67 813 Staphylococcus aureus (strain NCTC 8325 / PS 47)
68 804 Pectobacterium atrosepticum (strain SCRI 1043 / ATCC BAA-672)
69 796 Vibrio parahaemolyticus serotype O3:K6 (strain RIMD 2210633)
70 791 Escherichia coli (strain SMS-3-5 / SECEC)
71 788 Aquifex aeolicus (strain VF5)
72 779 Escherichia coli O127:H6 (strain E2348/69 / EPEC)
73 771 Escherichia coli (strain K12 / DH10B)
74 770 Pasteurella multocida (strain Pm70)
75 767 Escherichia coli O17:K52:H18 (strain UMN026 / ExPEC)
76 765 Escherichia coli (strain K12 / MC4100 / BW2952)
77 762 Escherichia coli (strain 55989 / EAEC)
78 761 Escherichia coli O8 (strain IAI1)
79 760 Shigella flexneri serotype 5b (strain 8401)
80 760 Staphylococcus epidermidis
81 760 Staphylococcus epidermidis (strain ATCC 12228 / FDA PCI 1200)
82 759 Escherichia coli O45:K1 (strain S88 / ExPEC)
83 758 Bacillus anthracis
84 756 Escherichia coli (strain SE11)
85 753 Escherichia coli O7:K1 (strain IAI39 / ExPEC)
86 749 Photorhabdus laumondii subsp. laumondii (strain DSM 15139 / CIP 105565 / TT01)
87 748 Escherichia coli O157:H7 (strain EC4115 / EHEC)
88 744 Halalkalibacterium halodurans
89 739 Yersinia enterocolitica serotype O:8 / biotype 1B (strain NCTC 13174 / 8081)
90 739 Escherichia coli
91 738 Pseudomonas putida
92 733 Vibrio vulnificus (strain CMCP6)
93 731 Escherichia coli O81 (strain ED1a)
94 722 Salmonella enteritidis PT4 (strain P125109)
95 719 Klebsiella pneumoniae subsp. pneumoniae (strain ATCC 700721 / MGH 78578)
96 718 Vibrio vulnificus (strain YJ016)
97 716 Salmonella paratyphi B (strain ATCC BAA-1250 / SPB7)
98 715 Enterobacter sp. (strain 638)
99 715 Escherichia coli O1:K1 / APEC
100 715 Yersinia pestis bv. Antiqua (strain Nepal516)
101 714 Salmonella paratyphi A (strain AKU_12601)
102 713 Yersinia pseudotuberculosis serotype O:1b (strain IP 31758)
103 713 Salmonella newport (strain SL254)
104 713 Salmonella agona (strain SL483)
105 712 Salmonella schwarzengrund (strain CVM19633)
106 711 Yersinia pestis bv. Antiqua (strain Antiqua)
107 710 Salmonella heidelberg (strain SL476)
108 708 Nostoc sp. (strain PCC 7120 / SAG 25.82 / UTEX 2576)
109 702 Salmonella dublin (strain CT_02021853)
110 699 Klebsiella variicola (strain 342) (Klebsiella pneumoniae)
111 698 Shigella boydii serotype 18 (strain CDC 3083-94 / BS512)
112 695 Escherichia fergusonii
113 692 Pan troglodytes (Chimpanzee)
114 686 Mycoplasma pneumoniae (strain ATCC 29342 / M129 / Subtype 1)
115 684 Salmonella gallinarum (strain 287/91 / NCTC 13346)
116 683 Pseudomonas syringae pv. tomato (strain ATCC BAA-871 / DC3000)
117 679 Agrobacterium fabrum (strain C58 / ATCC 33970) (Agrobacterium tumefaciens
118 679 Citrobacter koseri (strain ATCC BAA-895 / CDC 4225-83 / SGSC4696)
119 679 Staphylococcus aureus (strain USA300)
120 672 Serratia proteamaculans (strain 568)
121 672 Bacillus cereus
122 669 Mycobacterium leprae (strain TN)
123 667 Yarrowia lipolytica (strain CLIB 122 / E 150) (Yeast) (Candida lipolytica)
124 667 Bradyrhizobium diazoefficiens
125 667 Yersinia pestis (strain Pestoides F)
126 663 Shewanella oneidensis
127 658 Sinorhizobium fredii (strain NBRC 101917 / NGR234)
128 653 Debaryomyces hansenii
129 643 Staphylococcus aureus (strain bovine RF122 / ET3-1)
130 642 Yersinia pseudotuberculosis serotype O:3 (strain YPIII)
131 642 Salmonella arizonae (strain ATCC BAA-731 / CDC346-86 / RSK2980)
132 634 Yersinia pseudotuberculosis serotype IB (strain PB1/+)
133 623 Methanothermobacter thermautotrophicus
134 623 Cronobacter sakazakii (strain ATCC BAA-894) (Enterobacter sakazakii)
135 622 Treponema pallidum (strain Nichols)
136 622 Listeria monocytogenes serovar 1/2a (strain ATCC BAA-679 / EGD-e)
137 620 Pseudomonas aeruginosa (strain UCBPP-PA14)
138 615 Xanthomonas campestris pv. campestris
139 614 Mesorhizobium japonicum (Mesorhizobium loti
140 614 Staphylococcus haemolyticus (strain JCSC1435)
141 613 Helicobacter pylori (strain ATCC 700392 / 26695) (Campylobacter pylori)
142 611 Mycolicibacterium smegmatis (strain ATCC 700084 / mc(2)155)
143 605 Listeria innocua serovar 6a (strain ATCC BAA-680 / CLIP 11262)
144 604 Ralstonia nicotianae (strain ATCC BAA-1114 / GMI1000) (Ralstonia solanacearum)
145 602 Staphylococcus saprophyticus subsp. saprophyticus
146 602 Photobacterium profundum (strain SS9)
147 601 Salmonella paratyphi C (strain RKS4594)
148 600 Yersinia pestis bv. Antiqua (strain Angola)
149 595 Bacillus cereus (strain ATCC 10987 / NRS 248)
150 592 Neisseria meningitidis serogroup B (strain ATCC BAA-335 / MC58)
151 591 Pectobacterium carotovorum subsp. carotovorum (strain PC1)
152 588 Aspergillus oryzae (strain ATCC 42149 / RIB 40) (Yellow koji mold)
153 584 Rickettsia prowazekii (strain Madrid E)
154 582 Caenorhabditis briggsae
155 579 Brucella suis biovar 1 (strain 1330)
156 576 Brucella melitensis biotype 1
157 575 Caulobacter vibrioides (strain ATCC 19089 / CIP 103742 / CB 15)
158 573 Aliivibrio fischeri (strain ATCC 700601 / ES114) (Vibrio fischeri)
159 572 Buchnera aphidicola subsp. Acyrthosiphon pisum (strain APS)
160 569 Bacillus thuringiensis subsp. konkukian (strain 97-27)
161 568 Helicobacter pylori (strain J99 / ATCC 700824) (Campylobacter pylori J99)
162 568 Pseudomonas syringae pv. syringae (strain B728a)
163 567 Thermotoga maritima
164 566 Bacillus licheniformis
165 562 Bacillus cereus (strain ZK / E33L)
166 562 Buchnera aphidicola subsp. Schizaphis graminum (strain Sg)
167 561 Xanthomonas axonopodis pv. citri (strain 306)
168 559 Clostridium acetobutylicum
169 555 Pseudomonas fluorescens (strain Pf0-1)
170 554 Neisseria meningitidis serogroup A / serotype 4A (strain DSM 15465 / Z2491)
171 554 Pseudomonas fluorescens (strain ATCC BAA-477 / NRRL B-23932 / Pf-5)
172 553 Oceanobacillus iheyensis
173 547 Pseudomonas savastanoi pv. phaseolicola (Pseudomonas syringae pv. phaseolicola
174 543 Corynebacterium glutamicum
175 541 Lactococcus lactis subsp. lactis (strain IL1403) (Streptococcus lactis)
176 533 Bordetella bronchiseptica (strain ATCC BAA-588 / NCTC 13252 / RB50)
177 531 Erwinia tasmaniensis
178 530 Listeria monocytogenes serotype 4b (strain F2365)
179 529 Sodalis glossinidius (strain morsitans)
180 525 Staphylococcus aureus (strain Newman)
181 525 Deinococcus radiodurans
182 523 Vibrio cholerae serotype O1 (strain ATCC 39541 / Classical Ogawa 395 / O395)
183 522 Xylella fastidiosa (strain 9a5c)
184 519 Chromobacterium violaceum
185 519 Methanosarcina acetivorans (strain ATCC 35395 / DSM 2834 / JCM 12185 / C2A)
186 519 Streptococcus pneumoniae serotype 4 (strain ATCC BAA-334 / TIGR4)
187 516 Bordetella pertussis (strain Tohama I / ATCC BAA-589 / NCTC 13251)
188 515 Xylella fastidiosa (strain Temecula1 / ATCC 700964)
189 512 Pyrococcus furiosus (strain ATCC 43587 / DSM 3638 / JCM 8422 / Vc1)
190 512 Geobacillus kaustophilus (strain HTA426)
191 512 Haemophilus ducreyi (strain 35000HP / ATCC 700724)
192 512 Pseudomonas paraeruginosa (strain DSM 24068 / PA7) (Pseudomonas aeruginosa
193 511 Solanum lycopersicum (Tomato) (Lycopersicon esculentum)
194 511 Acinetobacter baylyi (strain ATCC 33305 / BD413 / ADP1)
195 511 Streptomyces avermitilis
196 509 Streptococcus pneumoniae (strain ATCC BAA-255 / R6)
197 508 Bordetella parapertussis (strain 12822 / ATCC BAA-587 / NCTC 13253)
198 507 Buchnera aphidicola subsp. Baizongia pistaciae (strain Bp)
199 506 Nicotiana tabacum (Common tobacco)
200 505 Saccharolobus solfataricus (strain ATCC 35092 / DSM 1617 / JCM 11322 / P2)
201 504 Pseudomonas entomophila (strain L48)
202 501 Methanosarcina mazei
203 501 Burkholderia pseudomallei (strain K96243)
204 499 Brucella abortus biovar 1 (strain 9-941)
205 499 Haemophilus influenzae (strain 86-028NP)
206 498 Thermosynechococcus vestitus (strain NIES-2133 / IAM M-273 / BP-1)
207 497 Xanthomonas campestris pv. campestris (strain 8004)
208 497 Synechococcus elongatus (strain ATCC 33912 / PCC 7942 / FACHB-805)
209 497 Pyrococcus horikoshii
210 497 Proteus mirabilis (strain HI4320)
211 496 Rickettsia conorii (strain ATCC VR-613 / Malish 7)
212 496 Shouchella clausii (strain KSM-K16) (Alkalihalobacillus clausii)
213 495 Halobacterium salinarum (strain ATCC 700922 / JCM 11081 / NRC-1)
214 493 Brucella abortus (strain 2308)
215 492 Bacillus velezensis (strain DSM 23117 / BGSC 10A6 / LMG 26770 / FZB42)
216 491 Vibrio campbellii (strain ATCC BAA-1116)
217 488 Shewanella sp. (strain MR-7)
218 486 Mannheimia succiniciproducens (strain KCTC 0769BP / MBEL55E)
219 485 Pseudomonas aeruginosa (strain LESB58)
220 485 Shewanella sp. (strain MR-4)
221 484 Staphylococcus aureus (strain Mu3 / ATCC 700698)
222 483 Mycoplasma genitalium (strain ATCC 33530 / DSM 19775 / NCTC 10195 / G37)
223 483 Lactiplantibacillus plantarum (strain ATCC BAA-793 / NCIMB 8826 / WCFS1)
224 480 Pseudomonas putida
225 478 Cupriavidus necator
226 478 Pyrococcus abyssi (strain GE5 / Orsay)
227 476 Enterococcus faecalis (strain ATCC 700802 / V583)
228 475 Burkholderia lata
229 475 Campylobacter jejuni subsp. jejuni serotype O:2
230 472 Rhodopseudomonas palustris (strain ATCC BAA-98 / CGA009)
231 471 Cereibacter sphaeroides
232 470 Clostridium perfringens (strain 13 / Type A)
233 468 Shewanella sp. (strain ANA-3)
234 468 Shewanella frigidimarina (strain NCIMB 400)
235 468 Pseudomonas putida (strain GB-1)
236 467 Aeromonas hydrophila subsp. hydrophila
237 466 Xanthomonas euvesicatoria pv. vesicatoria (strain 85-10)
238 465 Trichormus variabilis (strain ATCC 29413 / PCC 7937) (Anabaena variabilis)
239 463 Burkholderia mallei (strain ATCC 23344)
240 462 Cupriavidus pinatubonensis (strain JMP 134 / LMG 1197) (Cupriavidus necator
241 461 Ovis aries (Sheep)
242 460 Methylococcus capsulatus (strain ATCC 33009 / NCIMB 11132 / Bath)
243 457 Rickettsia felis (strain ATCC VR-1525 / URRWXCal2) (Rickettsia azadi)
244 455 Staphylococcus aureus (strain JH1)
245 455 Xanthomonas oryzae pv. oryzae (strain MAFF 311018)
246 455 Shewanella baltica (strain OS185)
247 453 Streptococcus mutans serotype c (strain ATCC 700610 / UA159)
248 453 Mycolicibacterium paratuberculosis (strain ATCC BAA-968 / K-10)
249 453 Pseudomonas putida (strain W619)
250 452 Caldanaerobacter subterraneus subsp. tengcongensis
2.3 Taxonomic distribution of the sequences
 Kingdom sequences (% of the database)
Archaea 19842 ( 3%)
Bacteria 336999 ( 59%)
Eukaryota 200289 ( 35%)
Viruses 17497 ( 3%)
Within Eukaryota:
Kingdom sequences (% of the database)
Archaea 19842 ( 3%)
Bacteria 336999 ( 59%)
Eukaryota 200289 ( 35%)
Viruses 17497 ( 3%)
Within Eukaryota:
 Category sequences (% of Eukaryota) (% of the complete database)
Human 20432 ( 10%) ( 4%)
Other Mammalia 47531 ( 24%) ( 8%)
Other Vertebrata 19057 ( 10%) ( 3%)
Viridiplantae 42002 ( 21%) ( 7%)
Fungi 38077 ( 19%) ( 7%)
Insecta 10104 ( 5%) ( 2%)
Nematoda 5418 ( 3%) ( 1%)
Other 17668 ( 9%) ( 3%)
3. SEQUENCE SIZE
Repartition of the sequences by size (excluding fragments)
From To Number From To Number
1- 50 10063 1001-1100 4181
51- 100 43847 1101-1200 2942
101- 150 60138 1201-1300 2236
151- 200 59834 1301-1400 2098
201- 250 58732 1401-1500 1707
251- 300 52750 1501-1600 849
301- 350 53196 1601-1700 653
351- 400 46288 1701-1800 606
401- 450 37946 1801-1900 543
451- 500 30836 1901-2000 406
501- 550 22592 2001-2100 283
551- 600 15985 2101-2200 399
601- 650 13259 2201-2300 348
651- 700 9486 2301-2400 243
701- 750 7942 2401-2500 202
751- 800 5755 >2500 1527
801- 850 4942
851- 900 5350
901- 950 4159
951-1000 3038
Category sequences (% of Eukaryota) (% of the complete database)
Human 20432 ( 10%) ( 4%)
Other Mammalia 47531 ( 24%) ( 8%)
Other Vertebrata 19057 ( 10%) ( 3%)
Viridiplantae 42002 ( 21%) ( 7%)
Fungi 38077 ( 19%) ( 7%)
Insecta 10104 ( 5%) ( 2%)
Nematoda 5418 ( 3%) ( 1%)
Other 17668 ( 9%) ( 3%)
3. SEQUENCE SIZE
Repartition of the sequences by size (excluding fragments)
From To Number From To Number
1- 50 10063 1001-1100 4181
51- 100 43847 1101-1200 2942
101- 150 60138 1201-1300 2236
151- 200 59834 1301-1400 2098
201- 250 58732 1401-1500 1707
251- 300 52750 1501-1600 849
301- 350 53196 1601-1700 653
351- 400 46288 1701-1800 606
401- 450 37946 1801-1900 543
451- 500 30836 1901-2000 406
501- 550 22592 2001-2100 283
551- 600 15985 2101-2200 399
601- 650 13259 2201-2300 348
651- 700 9486 2301-2400 243
701- 750 7942 2401-2500 202
751- 800 5755 >2500 1527
801- 850 4942
851- 900 5350
901- 950 4159
951-1000 3038
 The average sequence length in UniProtKB/Swiss-Prot is 362 amino acids.
The shortest sequence is GWA_SEPOF (P83570): 2 amino acids.
The longest sequence is TITIN_MOUSE (A2ASS6): 35213 amino acids.
4. JOURNAL CITATIONS
Note: the following citation statistics reflect the number of distinct
journal citations.
Total number of journals cited in this release of UniProtKB/Swiss-Prot: 3256
4.1 Table of the frequency of journal citations
Journals cited 1x: 1018
2x: 432
3x: 233
4x: 154
5x: 128
6x: 102
7x: 64
8x: 70
9x: 63
10x: 36
11- 20x: 257
21- 50x: 283
51-100x: 144
>100x: 272
4.2 List of the most cited journals in UniProtKB/Swiss-Prot
Nb Citations Journal name
-- --------- -------------------------------------------------------------
1 27952 Journal of Biological Chemistry
2 13174 Proceedings of the National Academy of Sciences of the U.S.A.
3 7396 Journal of Bacteriology
4 6197 Biochemical and Biophysical Research Communications
5 6014 Biochemistry
6 5511 Nucleic Acids Research
7 5415 Nature
8 5172 FEBS Letters
9 5115 The EMBO Journal
10 4911 Gene
11 4725 Journal of Molecular Biology
12 4667 Molecular and Cellular Biology
13 4121 Biochimica et Biophysica Acta
14 3985 Cell
15 3712 Journal of Virology
16 3538 European Journal of Biochemistry
17 3513 Science
18 3251 Biochemical Journal
19 3002 Molecular Microbiology
20 2844 Plant Physiology
21 2829 PLoS ONE
22 2549 Genomics
23 2501 The American Journal of Human Genetics
24 2432 Journal of Cell Biology
25 2217 The Plant Cell
26 2079 Human Molecular Genetics
27 2049 The Plant Journal
28 1994 Genes and Development
29 1976 Molecular Cell
30 1947 Virology
31 1929 Plant Molecular Biology
32 1875 Nature Genetics
33 1871 Molecular Biology of the Cell
34 1859 Development
35 1774 Journal of Immunology
36 1739 Nature Communications
37 1696 Human Mutation
38 1578 Oncogene
39 1519 Structure
40 1452 Journal of Biochemistry
41 1452 Genetics
42 1444 Molecular and General Genetics
43 1440 Journal of Cell Science
44 1327 Blood
45 1302 Infection and Immunity
46 1220 Microbiology
47 1212 Developmental Biology
48 1200 Journal of General Virology
49 1179 Current Biology
50 1172 Archives of Biochemistry and Biophysics
51 1099 Scientific Reports
52 1078 Journal of Neuroscience
53 1069 Applied and Environmental Microbiology
54 1013 Acta Crystallographica, Section D
55 973 PLoS Genetics
56 963 FEMS Microbiology Letters
57 940 Cancer Research
58 912 American Journal of Physiology
59 905 Toxicon
60 903 Protein Science
61 892 Journal of Clinical Investigation
62 860 Yeast
63 856 Neuron
64 807 The Journal of Experimental Medicine
65 783 PLoS Pathogens
66 779 Human Genetics
67 776 Plant and Cell Physiology
68 775 Nature Structural and Molecular Biology
69 745 Journal of Medical Genetics
70 740 The FEBS Journal
71 715 Proteins
72 684 Nature Cell Biology
73 682 Mechanisms of Development
74 661 Bioscience, Biotechnology, and Biochemistry
75 656 Nature Structural Biology
76 649 Antimicrobial Agents and Chemotherapy
77 618 Cell Reports
78 617 Developmental Cell
79 611 Current Genetics
80 584 Journal of Neurochemistry
81 564 Journal of the American Chemical Society
82 561 Molecular Endocrinology
83 558 The Journal of Clinical Endocrinology and Metabolism
84 555 Endocrinology
85 549 Molecular and Biochemical Parasitology
86 543
87 520 Eukaryotic Cell
88 505 Experimental Cell Research
89 504 EMBO Reports
90 499 RNA
91 495 Mammalian Genome
92 490 American Journal of Medical Genetics. Part A
93 486 The FASEB Journal
94 483 Peptides
95 477 Journal of Experimental Botany
96 462 Planta
97 449 Molecular Pharmacology
98 445 Acta Crystallographica, Section F
99 444 European Journal of Human Genetics
100 437 Clinical Genetics
101 435 Immunogenetics
102 432 Molecular Plant-Microbe Interactions
103 426 Immunity
104 426 Molecular Biology and Evolution
105 422 Journal of Investigative Dermatology
106 407 Journal of Molecular Evolution
107 402 Neurology
108 400 Biochimie
109 398 DNA and Cell Biology
110 387 Biology of Reproduction
111 381 Comparative Biochemistry and Physiology
112 381 DNA Sequence
113 376 PLoS Biology
114 372 Genes to Cells
115 368 Applied Microbiology and Biotechnology
116 366 Nature Immunology
117 366 Virus Research
118 359 Journal of Lipid Research
119 356 Journal of Medicinal Chemistry
120 352 BMC Genomics
121 350 Developmental Dynamics
122 346 The New England Journal of Medicine
123 343 Brain Research. Molecular Brain Research
124 341 Annals of Neurology
125 329 European Journal of Immunology
126 321 Journal of Human Genetics
127 321 Genome Research
128 315 Nature Chemical Biology
129 315 Investigative Ophthalmology and Visual Science
130 301 Brain
131 300 Glycobiology
132 299 Biological Chemistry Hoppe-Seyler
133 286 Fungal Genetics and Biology
134 285 Journal of General Microbiology
135 285 Archives of Microbiology
136 281 Cytogenetics and Cell Genetics
137 271 Traffic
138 271 Protein Expression and Purification
139 270 Cell Research
140 269 Molecular Genetics and Metabolism
141 266 Nature Medicine
142 263 Molecular Immunology
143 263 Phytochemistry
144 258 Journal of Cellular Biochemistry
145 254 Cell Cycle
146 246 Circulation Research
147 240 Insect Biochemistry and Molecular Biology
148 240 Diabetes
149 238 New Phytologist
150 237 Chemistry and Biology
5. STATISTICS FOR SOME LINE TYPES
The following table summarizes the total number of some UniProtKB/Swiss-Prot lines,
as well as the number of entries with at least one such line, and the
frequency of the lines.
Total Number of Average
Line type / subtype number entries per entry
------------------------------------ -------- --------- ---------
References (RL) 1338719 2.33
Journal 1167917 480978 2.03 1
Submitted to EMBL/GenBank/DDBJ 159128 143141 0.28 2
Submitted to other databases 7944 7239 0.01 3
Book citation 1877 1854 <0.01 4
Plant Gene Register 613 600 <0.01 5
Unpublished observations 543 539 <0.01 6
Thesis 477 474 <0.01 7
Patent 214 207 <0.01 8
Worm Breeder's Gazette 6 6 <0.01 9
Total number of distinct authors cited in UniProtKB/Swiss-Prot: 489697
Total Number of Average
Line type / subtype number entries per entry Rank
------------------------------------ -------- --------- --------- ----
Comments (CC) 2802336 4.88
ACTIVITY REGULATION 19539 19400 0.03 17
ALLERGEN 961 961 <0.01 26
ALTERNATIVE PRODUCTS 26020 26020 0.05 14
BIOPHYSICOCHEMICAL PROPERTIES 12354 12292 0.02 20
BIOTECHNOLOGY 2342 2281 <0.01 24
CATALYTIC ACTIVITY 357782 260363 0.62 4
CAUTION 14627 14324 0.03 19
COFACTOR 135473 123006 0.24 7
DEVELOPMENTAL STAGE 14912 14783 0.03 18
DISEASE 8698 5835 0.02 21
DISRUPTION PHENOTYPE 23450 23377 0.04 16
DOMAIN 62834 53157 0.11 9
FUNCTION 500305 473996 0.87 2
INDUCTION 27070 26950 0.05 13
INTERACTION 25638 25638 0.04 15
MASS SPECTROMETRY 7707 5971 0.01 22
MISCELLANEOUS 46679 41012 0.08 11
PATHWAY 145085 131041 0.25 6
PHARMACEUTICAL 169 162 <0.01 29
POLYMORPHISM 1515 1392 <0.01 25
PTM 67522 47537 0.12 8
RNA EDITING 646 646 <0.01 28
SEQUENCE CAUTION 45415 45345 0.08 12
SIMILARITY 522913 518545 0.91 1
SUBCELLULAR LOCATION 370136 361146 0.64 3
SUBUNIT 304021 298021 0.53 5
TISSUE SPECIFICITY 52275 51527 0.09 10
TOXIC DOSE 893 714 <0.01 27
WEB RESOURCE 5355 4823 0.01 23
Total number of comment topics: 29
Total Number of Average
Line type / subtype number entries per entry Rank
------------------------------------ -------- --------- --------- ----
Features (FT) 5659423 9.85
ACT_SITE 179698 107127 0.31 10
BINDING 1287551 222598 2.24 1
CARBOHYD 126559 32187 0.22 14
CHAIN 583134 566838 1.01 2
COILED 22877 15826 0.04 25
COMPBIAS 267854 93109 0.47 8
CONFLICT 140100 48791 0.24 13
CROSSLNK 25906 9283 0.05 24
DISULFID 141462 37439 0.25 12
DNA_BIND 12308 11027 0.02 31
DOMAIN 220216 134721 0.38 9
HELIX 374823 31767 0.65 5
INIT_MET 17661 17607 0.03 26
INTRAMEM 3194 1527 0.01 34
LIPID 14059 8959 0.02 28
MOD_RES 268774 75331 0.47 7
MOTIF 49441 31809 0.09 21
MUTAGEN 108262 21627 0.19 16
NON_CONS 2641 829 <0.01 35
NON_STD 360 285 <0.01 36
NON_TER 12570 9667 0.02 30
PEPTIDE 12886 8910 0.02 29
PROPEP 15633 13364 0.03 27
REGION 329961 151799 0.57 6
REPEAT 110205 15302 0.19 15
SIGNAL 45273 45272 0.08 22
SITE 69435 37261 0.12 19
STRAND 378775 29914 0.66 4
TOPO_DOM 154982 31064 0.27 11
TRANSIT 9674 9551 0.02 32
TRANSMEM 385580 80748 0.67 3
TURN 90810 26005 0.16 18
UNSURE 5773 909 0.01 33
VAR_SEQ 53476 22769 0.09 20
VARIANT 106543 17640 0.19 17
ZN_FING 30967 13240 0.05 23
Total number of feature keys: 36
Total Number of Average
Line type / subtype number entries per entry Rank Category
------------------------------------ -------- --------- --------- ---- -------------------------------------------
Cross-references (DR) 21823477 37.98
ABCD 3196 3196 0.01 123 Protocols and materials databases
Agora 18462 18413 0.03 86 Miscellaneous databases
AGR 69344 68617 0.12 43 Organism-specific databases
Allergome 2047 1316 <0.01 133 Protein family/group databases
AlphaFoldDB 549408 549408 0.96 10 3D structure databases
Antibodypedia 32354 32245 0.06 62 Protocols and materials databases
AntiFam 22 22 <0.01 169 Family and domain databases
ArachnoServer 1148 1138 <0.01 141 Organism-specific databases
Araport 16438 16342 0.03 92 Organism-specific databases
Bgee 61985 61984 0.11 45 Gene expression databases
BindingDB 6929 6929 0.01 109 Chemistry databases
BioCyc 48261 44210 0.08 55 Enzyme and pathway databases
BioGRID 62481 60445 0.11 44 Protein-protein interaction databases
BioGRID-ORCS 45145 44557 0.08 56 Miscellaneous databases
BioMuta 20286 20259 0.04 74 Genetic variation databases
BMRB 6914 6914 0.01 110 3D structure databases
BRENDA 20504 18683 0.04 71 Enzyme and pathway databases
CarbonylDB 1159 1159 <0.01 140 PTM databases
CARD 321 319 <0.01 158 Protein family/group databases
CAZy 9723 8755 0.02 100 Protein family/group databases
CCDS 49766 34842 0.09 52 Sequence databases
CD-CODE 10734 8219 0.02 98 Miscellaneous databases
CDD 393904 309774 0.69 17 Family and domain databases
CGD 2160 2143 <0.01 131 Organism-specific databases
ChEMBL 9290 9110 0.02 101 Chemistry databases
ChiTaRS 15285 15251 0.03 95 Miscellaneous databases
CIViC 569 568 <0.01 150 Organism-specific databases
ClinPGx 18028 18009 0.03 88 Organism-specific databases
CollecTF 138 138 <0.01 161 Gene expression databases
ComplexPortal 19302 9733 0.03 82 Protein-protein interaction databases
ConoServer 967 879 <0.01 144 Organism-specific databases
CORUM 8090 8090 0.01 105 Protein-protein interaction databases
CPTAC 3472 1929 0.01 119 Proteomic databases
CPTC 410 410 <0.01 153 Protocols and materials databases
CTD 77698 77047 0.14 41 Organism-specific databases
DEPOD 254 254 <0.01 160 PTM databases
dictyBase 4228 4114 0.01 116 Organism-specific databases
DIP 17580 17539 0.03 90 Protein-protein interaction databases
DisGeNET 17613 17415 0.03 89 Organism-specific databases
DisProt 2825 2800 <0.01 126 Family and domain databases
DMDM 16164 16163 0.03 94 Genetic variation databases
DNASU 48556 48477 0.08 54 Protocols and materials databases
DrugBank 35601 4939 0.06 61 Chemistry databases
DrugCentral 2982 2982 0.01 125 Chemistry databases
EchoBASE 4158 4158 0.01 117 Organism-specific databases
eggNOG 340513 334622 0.59 20 Phylogenomic databases
ELM 1815 1815 <0.01 134 Protein-protein interaction databases
EMBL 1011147 561609 1.76 3 Sequence databases
EMDB 127943 11682 0.22 32 3D structure databases
Ensembl 122986 51864 0.21 34 Genome annotation databases
EnsemblBacteria 55578 55400 0.10 48 Genome annotation databases
EnsemblFungi 19513 19196 0.03 79 Genome annotation databases
EnsemblMetazoa 19685 12907 0.03 78 Genome annotation databases
EnsemblPlants 26391 6359 0.05 66 Genome annotation databases
EnsemblProtists 1740 1593 <0.01 135 Genome annotation databases
ESTHER 3035 3032 0.01 124 Protein family/group databases
euHCVdb 55 44 <0.01 166 Organism-specific databases
EvolutionaryTrace 22703 22703 0.04 69 Miscellaneous databases
ExpressionAtlas 51312 51312 0.09 50 Gene expression databases
FlyBase 3997 3888 0.01 118 Organism-specific databases
FunCoup 143520 143520 0.25 30 Protein-protein interaction databases
FunFam 558753 327653 0.97 9 Family and domain databases
Gene3D 800342 478318 1.39 6 Family and domain databases
GeneCards 20377 20247 0.04 73 Organism-specific databases
GeneID 315366 288167 0.55 23 Genome annotation databases
GeneReviews 1623 1619 <0.01 136 Organism-specific databases
GeneTree 48938 48921 0.09 53 Phylogenomic databases
GeneWiki 10351 10269 0.02 99 Miscellaneous databases
GenomeRNAi 22327 22326 0.04 70 Miscellaneous databases
GlyConnect 2372 2215 <0.01 128 PTM databases
GlyCosmos 28908 28908 0.05 64 PTM databases
GlyGen 39091 39091 0.07 59 PTM databases
GO 3358100 553982 5.84 1 Ontologies
Gramene 51809 22301 0.09 49 Genome annotation databases
GuidetoPHARMACOLOGY 2299 2299 <0.01 130 Chemistry databases
HAMAP 331036 328099 0.58 22 Family and domain databases
HGNC 20382 20256 0.04 72 Organism-specific databases
HOGENOM 428629 428629 0.75 16 Phylogenomic databases
HPA 19354 19215 0.03 81 Organism-specific databases
IDEAL 1101 1101 <0.01 142 Family and domain databases
IMGT_GENE-DB 267 267 <0.01 159 Protein family/group databases
InParanoid 164709 164709 0.29 26 Phylogenomic databases
IntAct 57960 57960 0.10 46 Protein-protein interaction databases
InterPro 2597857 556937 4.52 2 Family and domain databases
iPTMnet 56791 56791 0.10 47 PTM databases
JaponicusDB 43 43 <0.01 167 Organism-specific databases
jPOST 29053 29053 0.05 63 Proteomic databases
KEGG 520948 484511 0.91 13 Genome annotation databases
LegioList 765 763 <0.01 147 Organism-specific databases
Leproma 672 669 <0.01 148 Organism-specific databases
MaizeGDB 529 525 <0.01 151 Organism-specific databases
MalaCards 7396 7383 0.01 107 Organism-specific databases
MANE-Select 18592 18480 0.03 85 Genome annotation databases
MassIVE 19140 19140 0.03 83 Proteomic databases
MEROPS 14264 13845 0.02 97 Protein family/group databases
MetOSite 3455 3455 0.01 120 PTM databases
MGI 17168 17125 0.03 91 Organism-specific databases
MIM 24144 16578 0.04 68 Organism-specific databases
MINT 24151 24151 0.04 67 Protein-protein interaction databases
MoonDB 348 348 <0.01 157 Protein family/group databases
MoonProt 368 368 <0.01 155 Protein family/group databases
NCBIfam 547439 347231 0.95 11 Family and domain databases
NIAGADS 76 76 <0.01 163 Organism-specific databases
OGP 373 373 <0.01 154 2D gel databases
OMA 120955 120955 0.21 35 Phylogenomic databases
OpenTargets 18614 18469 0.03 84 Organism-specific databases
Orphanet 8157 4441 0.01 104 Organism-specific databases
OrthoDB 270718 270718 0.47 24 Phylogenomic databases
PAN-GO 20210 20210 0.04 75 Phylogenomic databases
PANTHER 964680 505734 1.68 4 Family and domain databases
PathwayCommons 19433 19433 0.03 80 Enzyme and pathway databases
PATRIC 93326 93326 0.16 39 Genome annotation databases
PaxDb 154216 154216 0.27 27 Proteomic databases
PCDDB 134 134 <0.01 162 3D structure databases
PDB 361254 38379 0.63 19 3D structure databases
PDBsum 361254 38379 0.63 18 3D structure databases
PeptideAtlas 38922 38922 0.07 60 Proteomic databases
PeroxiBase 794 773 <0.01 146 Protein family/group databases
Pfam 871800 547272 1.52 5 Family and domain databases
Pharos 20192 20192 0.04 76 Miscellaneous databases
PHI-base 2470 1949 <0.01 127 Miscellaneous databases
PhosphoSitePlus 42259 42259 0.07 58 PTM databases
PhylomeDB 115823 115823 0.20 36 Phylogenomic databases
PIR 125284 114939 0.22 33 Sequence databases
PIRSF 111097 109925 0.19 37 Family and domain databases
PlantReactome 1436 824 <0.01 138 Enzyme and pathway databases
PomBase 5135 5131 0.01 114 Organism-specific databases
PRIDE 637 637 <0.01 149 Proteomic databases
PRINTS 151585 130114 0.26 28 Family and domain databases
PRO 100156 100156 0.17 38 Miscellaneous databases
ProMEX 489 489 <0.01 152 Proteomic databases
PROSITE 496275 313384 0.86 14 Family and domain databases
Proteomes 491388 450492 0.86 15 Miscellaneous databases
ProteomicsDB 72937 45501 0.13 42 Proteomic databases
PseudoCAP 2054 2054 <0.01 132 Organism-specific databases
Pumba 18203 18203 0.03 87 Proteomic databases
Reactome 148027 39443 0.26 29 Enzyme and pathway databases
REBASE 802 395 <0.01 145 Protein family/group databases
RefSeq 566363 442362 0.99 8 Sequence databases
REPRODUCTION-2DPAGE 1260 1039 <0.01 139 2D gel databases
RGD 8159 8158 0.01 103 Organism-specific databases
RNAct 43120 43120 0.08 57 Miscellaneous databases
SABIO-RK 5951 5951 0.01 112 Enzyme and pathway databases
SASBDB 1025 1025 <0.01 143 3D structure databases
SFLD 27770 9138 0.05 65 Family and domain databases
SGD 6753 6748 0.01 111 Organism-specific databases
SignaLink 19948 19948 0.03 77 Enzyme and pathway databases
SIGNOR 7769 7769 0.01 106 Enzyme and pathway databases
SMART 207127 149377 0.36 25 Family and domain databases
SMR 525917 525917 0.92 12 3D structure databases
STRENDA-DB 59 45 <0.01 164 Enzyme and pathway databases
STRING 337189 337189 0.59 21 Protein-protein interaction databases
SUPFAM 652113 461938 1.13 7 Family and domain databases
SwissLipids 1478 1394 <0.01 137 Chemistry databases
SwissPalm 14331 14331 0.02 96 PTM databases
TAIR 16432 16342 0.03 93 Organism-specific databases
TCDB 8821 8723 0.02 102 Protein family/group databases
TopDownProteomics 3236 2957 0.01 122 Proteomic databases
TubercuList 2359 2323 <0.01 129 Organism-specific databases
UCSC 51124 46623 0.09 51 Genome annotation databases
UniLectin 367 367 <0.01 156 Protein family/group databases
UniPathway 140553 126895 0.24 31 Enzyme and pathway databases
VEuPathDB 87783 80328 0.15 40 Organism-specific databases
VGNC 5155 5143 0.01 113 Organism-specific databases
WBParaSite 56 54 <0.01 165 Genome annotation databases
WormBase 6951 5108 0.01 108 Organism-specific databases
Xenbase 4758 4758 0.01 115 Organism-specific databases
YCharOS 36 36 <0.01 168 Protocols and materials databases
ZFIN 3298 3295 0.01 121 Organism-specific databases
Total number of cross-referenced databases: 169
6. AMINO ACID COMPOSITION
6.1 Composition in percent for the complete database
Ala (A) 8.25 Gln (Q) 3.93 Leu (L) 9.64 Ser (S) 6.66
Arg (R) 5.52 Glu (E) 6.71 Lys (K) 5.79 Thr (T) 5.36
Asn (N) 4.06 Gly (G) 7.07 Met (M) 2.41 Trp (W) 1.10
Asp (D) 5.46 His (H) 2.27 Phe (F) 3.86 Tyr (Y) 2.92
Cys (C) 1.38 Ile (I) 5.90 Pro (P) 4.74 Val (V) 6.85
Asx (B) 0.000 Glx (Z) 0.000 Xaa (X) 0.00
The average sequence length in UniProtKB/Swiss-Prot is 362 amino acids.
The shortest sequence is GWA_SEPOF (P83570): 2 amino acids.
The longest sequence is TITIN_MOUSE (A2ASS6): 35213 amino acids.
4. JOURNAL CITATIONS
Note: the following citation statistics reflect the number of distinct
journal citations.
Total number of journals cited in this release of UniProtKB/Swiss-Prot: 3256
4.1 Table of the frequency of journal citations
Journals cited 1x: 1018
2x: 432
3x: 233
4x: 154
5x: 128
6x: 102
7x: 64
8x: 70
9x: 63
10x: 36
11- 20x: 257
21- 50x: 283
51-100x: 144
>100x: 272
4.2 List of the most cited journals in UniProtKB/Swiss-Prot
Nb Citations Journal name
-- --------- -------------------------------------------------------------
1 27952 Journal of Biological Chemistry
2 13174 Proceedings of the National Academy of Sciences of the U.S.A.
3 7396 Journal of Bacteriology
4 6197 Biochemical and Biophysical Research Communications
5 6014 Biochemistry
6 5511 Nucleic Acids Research
7 5415 Nature
8 5172 FEBS Letters
9 5115 The EMBO Journal
10 4911 Gene
11 4725 Journal of Molecular Biology
12 4667 Molecular and Cellular Biology
13 4121 Biochimica et Biophysica Acta
14 3985 Cell
15 3712 Journal of Virology
16 3538 European Journal of Biochemistry
17 3513 Science
18 3251 Biochemical Journal
19 3002 Molecular Microbiology
20 2844 Plant Physiology
21 2829 PLoS ONE
22 2549 Genomics
23 2501 The American Journal of Human Genetics
24 2432 Journal of Cell Biology
25 2217 The Plant Cell
26 2079 Human Molecular Genetics
27 2049 The Plant Journal
28 1994 Genes and Development
29 1976 Molecular Cell
30 1947 Virology
31 1929 Plant Molecular Biology
32 1875 Nature Genetics
33 1871 Molecular Biology of the Cell
34 1859 Development
35 1774 Journal of Immunology
36 1739 Nature Communications
37 1696 Human Mutation
38 1578 Oncogene
39 1519 Structure
40 1452 Journal of Biochemistry
41 1452 Genetics
42 1444 Molecular and General Genetics
43 1440 Journal of Cell Science
44 1327 Blood
45 1302 Infection and Immunity
46 1220 Microbiology
47 1212 Developmental Biology
48 1200 Journal of General Virology
49 1179 Current Biology
50 1172 Archives of Biochemistry and Biophysics
51 1099 Scientific Reports
52 1078 Journal of Neuroscience
53 1069 Applied and Environmental Microbiology
54 1013 Acta Crystallographica, Section D
55 973 PLoS Genetics
56 963 FEMS Microbiology Letters
57 940 Cancer Research
58 912 American Journal of Physiology
59 905 Toxicon
60 903 Protein Science
61 892 Journal of Clinical Investigation
62 860 Yeast
63 856 Neuron
64 807 The Journal of Experimental Medicine
65 783 PLoS Pathogens
66 779 Human Genetics
67 776 Plant and Cell Physiology
68 775 Nature Structural and Molecular Biology
69 745 Journal of Medical Genetics
70 740 The FEBS Journal
71 715 Proteins
72 684 Nature Cell Biology
73 682 Mechanisms of Development
74 661 Bioscience, Biotechnology, and Biochemistry
75 656 Nature Structural Biology
76 649 Antimicrobial Agents and Chemotherapy
77 618 Cell Reports
78 617 Developmental Cell
79 611 Current Genetics
80 584 Journal of Neurochemistry
81 564 Journal of the American Chemical Society
82 561 Molecular Endocrinology
83 558 The Journal of Clinical Endocrinology and Metabolism
84 555 Endocrinology
85 549 Molecular and Biochemical Parasitology
86 543
87 520 Eukaryotic Cell
88 505 Experimental Cell Research
89 504 EMBO Reports
90 499 RNA
91 495 Mammalian Genome
92 490 American Journal of Medical Genetics. Part A
93 486 The FASEB Journal
94 483 Peptides
95 477 Journal of Experimental Botany
96 462 Planta
97 449 Molecular Pharmacology
98 445 Acta Crystallographica, Section F
99 444 European Journal of Human Genetics
100 437 Clinical Genetics
101 435 Immunogenetics
102 432 Molecular Plant-Microbe Interactions
103 426 Immunity
104 426 Molecular Biology and Evolution
105 422 Journal of Investigative Dermatology
106 407 Journal of Molecular Evolution
107 402 Neurology
108 400 Biochimie
109 398 DNA and Cell Biology
110 387 Biology of Reproduction
111 381 Comparative Biochemistry and Physiology
112 381 DNA Sequence
113 376 PLoS Biology
114 372 Genes to Cells
115 368 Applied Microbiology and Biotechnology
116 366 Nature Immunology
117 366 Virus Research
118 359 Journal of Lipid Research
119 356 Journal of Medicinal Chemistry
120 352 BMC Genomics
121 350 Developmental Dynamics
122 346 The New England Journal of Medicine
123 343 Brain Research. Molecular Brain Research
124 341 Annals of Neurology
125 329 European Journal of Immunology
126 321 Journal of Human Genetics
127 321 Genome Research
128 315 Nature Chemical Biology
129 315 Investigative Ophthalmology and Visual Science
130 301 Brain
131 300 Glycobiology
132 299 Biological Chemistry Hoppe-Seyler
133 286 Fungal Genetics and Biology
134 285 Journal of General Microbiology
135 285 Archives of Microbiology
136 281 Cytogenetics and Cell Genetics
137 271 Traffic
138 271 Protein Expression and Purification
139 270 Cell Research
140 269 Molecular Genetics and Metabolism
141 266 Nature Medicine
142 263 Molecular Immunology
143 263 Phytochemistry
144 258 Journal of Cellular Biochemistry
145 254 Cell Cycle
146 246 Circulation Research
147 240 Insect Biochemistry and Molecular Biology
148 240 Diabetes
149 238 New Phytologist
150 237 Chemistry and Biology
5. STATISTICS FOR SOME LINE TYPES
The following table summarizes the total number of some UniProtKB/Swiss-Prot lines,
as well as the number of entries with at least one such line, and the
frequency of the lines.
Total Number of Average
Line type / subtype number entries per entry
------------------------------------ -------- --------- ---------
References (RL) 1338719 2.33
Journal 1167917 480978 2.03 1
Submitted to EMBL/GenBank/DDBJ 159128 143141 0.28 2
Submitted to other databases 7944 7239 0.01 3
Book citation 1877 1854 <0.01 4
Plant Gene Register 613 600 <0.01 5
Unpublished observations 543 539 <0.01 6
Thesis 477 474 <0.01 7
Patent 214 207 <0.01 8
Worm Breeder's Gazette 6 6 <0.01 9
Total number of distinct authors cited in UniProtKB/Swiss-Prot: 489697
Total Number of Average
Line type / subtype number entries per entry Rank
------------------------------------ -------- --------- --------- ----
Comments (CC) 2802336 4.88
ACTIVITY REGULATION 19539 19400 0.03 17
ALLERGEN 961 961 <0.01 26
ALTERNATIVE PRODUCTS 26020 26020 0.05 14
BIOPHYSICOCHEMICAL PROPERTIES 12354 12292 0.02 20
BIOTECHNOLOGY 2342 2281 <0.01 24
CATALYTIC ACTIVITY 357782 260363 0.62 4
CAUTION 14627 14324 0.03 19
COFACTOR 135473 123006 0.24 7
DEVELOPMENTAL STAGE 14912 14783 0.03 18
DISEASE 8698 5835 0.02 21
DISRUPTION PHENOTYPE 23450 23377 0.04 16
DOMAIN 62834 53157 0.11 9
FUNCTION 500305 473996 0.87 2
INDUCTION 27070 26950 0.05 13
INTERACTION 25638 25638 0.04 15
MASS SPECTROMETRY 7707 5971 0.01 22
MISCELLANEOUS 46679 41012 0.08 11
PATHWAY 145085 131041 0.25 6
PHARMACEUTICAL 169 162 <0.01 29
POLYMORPHISM 1515 1392 <0.01 25
PTM 67522 47537 0.12 8
RNA EDITING 646 646 <0.01 28
SEQUENCE CAUTION 45415 45345 0.08 12
SIMILARITY 522913 518545 0.91 1
SUBCELLULAR LOCATION 370136 361146 0.64 3
SUBUNIT 304021 298021 0.53 5
TISSUE SPECIFICITY 52275 51527 0.09 10
TOXIC DOSE 893 714 <0.01 27
WEB RESOURCE 5355 4823 0.01 23
Total number of comment topics: 29
Total Number of Average
Line type / subtype number entries per entry Rank
------------------------------------ -------- --------- --------- ----
Features (FT) 5659423 9.85
ACT_SITE 179698 107127 0.31 10
BINDING 1287551 222598 2.24 1
CARBOHYD 126559 32187 0.22 14
CHAIN 583134 566838 1.01 2
COILED 22877 15826 0.04 25
COMPBIAS 267854 93109 0.47 8
CONFLICT 140100 48791 0.24 13
CROSSLNK 25906 9283 0.05 24
DISULFID 141462 37439 0.25 12
DNA_BIND 12308 11027 0.02 31
DOMAIN 220216 134721 0.38 9
HELIX 374823 31767 0.65 5
INIT_MET 17661 17607 0.03 26
INTRAMEM 3194 1527 0.01 34
LIPID 14059 8959 0.02 28
MOD_RES 268774 75331 0.47 7
MOTIF 49441 31809 0.09 21
MUTAGEN 108262 21627 0.19 16
NON_CONS 2641 829 <0.01 35
NON_STD 360 285 <0.01 36
NON_TER 12570 9667 0.02 30
PEPTIDE 12886 8910 0.02 29
PROPEP 15633 13364 0.03 27
REGION 329961 151799 0.57 6
REPEAT 110205 15302 0.19 15
SIGNAL 45273 45272 0.08 22
SITE 69435 37261 0.12 19
STRAND 378775 29914 0.66 4
TOPO_DOM 154982 31064 0.27 11
TRANSIT 9674 9551 0.02 32
TRANSMEM 385580 80748 0.67 3
TURN 90810 26005 0.16 18
UNSURE 5773 909 0.01 33
VAR_SEQ 53476 22769 0.09 20
VARIANT 106543 17640 0.19 17
ZN_FING 30967 13240 0.05 23
Total number of feature keys: 36
Total Number of Average
Line type / subtype number entries per entry Rank Category
------------------------------------ -------- --------- --------- ---- -------------------------------------------
Cross-references (DR) 21823477 37.98
ABCD 3196 3196 0.01 123 Protocols and materials databases
Agora 18462 18413 0.03 86 Miscellaneous databases
AGR 69344 68617 0.12 43 Organism-specific databases
Allergome 2047 1316 <0.01 133 Protein family/group databases
AlphaFoldDB 549408 549408 0.96 10 3D structure databases
Antibodypedia 32354 32245 0.06 62 Protocols and materials databases
AntiFam 22 22 <0.01 169 Family and domain databases
ArachnoServer 1148 1138 <0.01 141 Organism-specific databases
Araport 16438 16342 0.03 92 Organism-specific databases
Bgee 61985 61984 0.11 45 Gene expression databases
BindingDB 6929 6929 0.01 109 Chemistry databases
BioCyc 48261 44210 0.08 55 Enzyme and pathway databases
BioGRID 62481 60445 0.11 44 Protein-protein interaction databases
BioGRID-ORCS 45145 44557 0.08 56 Miscellaneous databases
BioMuta 20286 20259 0.04 74 Genetic variation databases
BMRB 6914 6914 0.01 110 3D structure databases
BRENDA 20504 18683 0.04 71 Enzyme and pathway databases
CarbonylDB 1159 1159 <0.01 140 PTM databases
CARD 321 319 <0.01 158 Protein family/group databases
CAZy 9723 8755 0.02 100 Protein family/group databases
CCDS 49766 34842 0.09 52 Sequence databases
CD-CODE 10734 8219 0.02 98 Miscellaneous databases
CDD 393904 309774 0.69 17 Family and domain databases
CGD 2160 2143 <0.01 131 Organism-specific databases
ChEMBL 9290 9110 0.02 101 Chemistry databases
ChiTaRS 15285 15251 0.03 95 Miscellaneous databases
CIViC 569 568 <0.01 150 Organism-specific databases
ClinPGx 18028 18009 0.03 88 Organism-specific databases
CollecTF 138 138 <0.01 161 Gene expression databases
ComplexPortal 19302 9733 0.03 82 Protein-protein interaction databases
ConoServer 967 879 <0.01 144 Organism-specific databases
CORUM 8090 8090 0.01 105 Protein-protein interaction databases
CPTAC 3472 1929 0.01 119 Proteomic databases
CPTC 410 410 <0.01 153 Protocols and materials databases
CTD 77698 77047 0.14 41 Organism-specific databases
DEPOD 254 254 <0.01 160 PTM databases
dictyBase 4228 4114 0.01 116 Organism-specific databases
DIP 17580 17539 0.03 90 Protein-protein interaction databases
DisGeNET 17613 17415 0.03 89 Organism-specific databases
DisProt 2825 2800 <0.01 126 Family and domain databases
DMDM 16164 16163 0.03 94 Genetic variation databases
DNASU 48556 48477 0.08 54 Protocols and materials databases
DrugBank 35601 4939 0.06 61 Chemistry databases
DrugCentral 2982 2982 0.01 125 Chemistry databases
EchoBASE 4158 4158 0.01 117 Organism-specific databases
eggNOG 340513 334622 0.59 20 Phylogenomic databases
ELM 1815 1815 <0.01 134 Protein-protein interaction databases
EMBL 1011147 561609 1.76 3 Sequence databases
EMDB 127943 11682 0.22 32 3D structure databases
Ensembl 122986 51864 0.21 34 Genome annotation databases
EnsemblBacteria 55578 55400 0.10 48 Genome annotation databases
EnsemblFungi 19513 19196 0.03 79 Genome annotation databases
EnsemblMetazoa 19685 12907 0.03 78 Genome annotation databases
EnsemblPlants 26391 6359 0.05 66 Genome annotation databases
EnsemblProtists 1740 1593 <0.01 135 Genome annotation databases
ESTHER 3035 3032 0.01 124 Protein family/group databases
euHCVdb 55 44 <0.01 166 Organism-specific databases
EvolutionaryTrace 22703 22703 0.04 69 Miscellaneous databases
ExpressionAtlas 51312 51312 0.09 50 Gene expression databases
FlyBase 3997 3888 0.01 118 Organism-specific databases
FunCoup 143520 143520 0.25 30 Protein-protein interaction databases
FunFam 558753 327653 0.97 9 Family and domain databases
Gene3D 800342 478318 1.39 6 Family and domain databases
GeneCards 20377 20247 0.04 73 Organism-specific databases
GeneID 315366 288167 0.55 23 Genome annotation databases
GeneReviews 1623 1619 <0.01 136 Organism-specific databases
GeneTree 48938 48921 0.09 53 Phylogenomic databases
GeneWiki 10351 10269 0.02 99 Miscellaneous databases
GenomeRNAi 22327 22326 0.04 70 Miscellaneous databases
GlyConnect 2372 2215 <0.01 128 PTM databases
GlyCosmos 28908 28908 0.05 64 PTM databases
GlyGen 39091 39091 0.07 59 PTM databases
GO 3358100 553982 5.84 1 Ontologies
Gramene 51809 22301 0.09 49 Genome annotation databases
GuidetoPHARMACOLOGY 2299 2299 <0.01 130 Chemistry databases
HAMAP 331036 328099 0.58 22 Family and domain databases
HGNC 20382 20256 0.04 72 Organism-specific databases
HOGENOM 428629 428629 0.75 16 Phylogenomic databases
HPA 19354 19215 0.03 81 Organism-specific databases
IDEAL 1101 1101 <0.01 142 Family and domain databases
IMGT_GENE-DB 267 267 <0.01 159 Protein family/group databases
InParanoid 164709 164709 0.29 26 Phylogenomic databases
IntAct 57960 57960 0.10 46 Protein-protein interaction databases
InterPro 2597857 556937 4.52 2 Family and domain databases
iPTMnet 56791 56791 0.10 47 PTM databases
JaponicusDB 43 43 <0.01 167 Organism-specific databases
jPOST 29053 29053 0.05 63 Proteomic databases
KEGG 520948 484511 0.91 13 Genome annotation databases
LegioList 765 763 <0.01 147 Organism-specific databases
Leproma 672 669 <0.01 148 Organism-specific databases
MaizeGDB 529 525 <0.01 151 Organism-specific databases
MalaCards 7396 7383 0.01 107 Organism-specific databases
MANE-Select 18592 18480 0.03 85 Genome annotation databases
MassIVE 19140 19140 0.03 83 Proteomic databases
MEROPS 14264 13845 0.02 97 Protein family/group databases
MetOSite 3455 3455 0.01 120 PTM databases
MGI 17168 17125 0.03 91 Organism-specific databases
MIM 24144 16578 0.04 68 Organism-specific databases
MINT 24151 24151 0.04 67 Protein-protein interaction databases
MoonDB 348 348 <0.01 157 Protein family/group databases
MoonProt 368 368 <0.01 155 Protein family/group databases
NCBIfam 547439 347231 0.95 11 Family and domain databases
NIAGADS 76 76 <0.01 163 Organism-specific databases
OGP 373 373 <0.01 154 2D gel databases
OMA 120955 120955 0.21 35 Phylogenomic databases
OpenTargets 18614 18469 0.03 84 Organism-specific databases
Orphanet 8157 4441 0.01 104 Organism-specific databases
OrthoDB 270718 270718 0.47 24 Phylogenomic databases
PAN-GO 20210 20210 0.04 75 Phylogenomic databases
PANTHER 964680 505734 1.68 4 Family and domain databases
PathwayCommons 19433 19433 0.03 80 Enzyme and pathway databases
PATRIC 93326 93326 0.16 39 Genome annotation databases
PaxDb 154216 154216 0.27 27 Proteomic databases
PCDDB 134 134 <0.01 162 3D structure databases
PDB 361254 38379 0.63 19 3D structure databases
PDBsum 361254 38379 0.63 18 3D structure databases
PeptideAtlas 38922 38922 0.07 60 Proteomic databases
PeroxiBase 794 773 <0.01 146 Protein family/group databases
Pfam 871800 547272 1.52 5 Family and domain databases
Pharos 20192 20192 0.04 76 Miscellaneous databases
PHI-base 2470 1949 <0.01 127 Miscellaneous databases
PhosphoSitePlus 42259 42259 0.07 58 PTM databases
PhylomeDB 115823 115823 0.20 36 Phylogenomic databases
PIR 125284 114939 0.22 33 Sequence databases
PIRSF 111097 109925 0.19 37 Family and domain databases
PlantReactome 1436 824 <0.01 138 Enzyme and pathway databases
PomBase 5135 5131 0.01 114 Organism-specific databases
PRIDE 637 637 <0.01 149 Proteomic databases
PRINTS 151585 130114 0.26 28 Family and domain databases
PRO 100156 100156 0.17 38 Miscellaneous databases
ProMEX 489 489 <0.01 152 Proteomic databases
PROSITE 496275 313384 0.86 14 Family and domain databases
Proteomes 491388 450492 0.86 15 Miscellaneous databases
ProteomicsDB 72937 45501 0.13 42 Proteomic databases
PseudoCAP 2054 2054 <0.01 132 Organism-specific databases
Pumba 18203 18203 0.03 87 Proteomic databases
Reactome 148027 39443 0.26 29 Enzyme and pathway databases
REBASE 802 395 <0.01 145 Protein family/group databases
RefSeq 566363 442362 0.99 8 Sequence databases
REPRODUCTION-2DPAGE 1260 1039 <0.01 139 2D gel databases
RGD 8159 8158 0.01 103 Organism-specific databases
RNAct 43120 43120 0.08 57 Miscellaneous databases
SABIO-RK 5951 5951 0.01 112 Enzyme and pathway databases
SASBDB 1025 1025 <0.01 143 3D structure databases
SFLD 27770 9138 0.05 65 Family and domain databases
SGD 6753 6748 0.01 111 Organism-specific databases
SignaLink 19948 19948 0.03 77 Enzyme and pathway databases
SIGNOR 7769 7769 0.01 106 Enzyme and pathway databases
SMART 207127 149377 0.36 25 Family and domain databases
SMR 525917 525917 0.92 12 3D structure databases
STRENDA-DB 59 45 <0.01 164 Enzyme and pathway databases
STRING 337189 337189 0.59 21 Protein-protein interaction databases
SUPFAM 652113 461938 1.13 7 Family and domain databases
SwissLipids 1478 1394 <0.01 137 Chemistry databases
SwissPalm 14331 14331 0.02 96 PTM databases
TAIR 16432 16342 0.03 93 Organism-specific databases
TCDB 8821 8723 0.02 102 Protein family/group databases
TopDownProteomics 3236 2957 0.01 122 Proteomic databases
TubercuList 2359 2323 <0.01 129 Organism-specific databases
UCSC 51124 46623 0.09 51 Genome annotation databases
UniLectin 367 367 <0.01 156 Protein family/group databases
UniPathway 140553 126895 0.24 31 Enzyme and pathway databases
VEuPathDB 87783 80328 0.15 40 Organism-specific databases
VGNC 5155 5143 0.01 113 Organism-specific databases
WBParaSite 56 54 <0.01 165 Genome annotation databases
WormBase 6951 5108 0.01 108 Organism-specific databases
Xenbase 4758 4758 0.01 115 Organism-specific databases
YCharOS 36 36 <0.01 168 Protocols and materials databases
ZFIN 3298 3295 0.01 121 Organism-specific databases
Total number of cross-referenced databases: 169
6. AMINO ACID COMPOSITION
6.1 Composition in percent for the complete database
Ala (A) 8.25 Gln (Q) 3.93 Leu (L) 9.64 Ser (S) 6.66
Arg (R) 5.52 Glu (E) 6.71 Lys (K) 5.79 Thr (T) 5.36
Asn (N) 4.06 Gly (G) 7.07 Met (M) 2.41 Trp (W) 1.10
Asp (D) 5.46 His (H) 2.27 Phe (F) 3.86 Tyr (Y) 2.92
Cys (C) 1.38 Ile (I) 5.90 Pro (P) 4.74 Val (V) 6.85
Asx (B) 0.000 Glx (Z) 0.000 Xaa (X) 0.00
 Legend: gray = aliphatic, red = acidic, green = small hydroxy,
blue = basic, black = aromatic, white = amide, yellow = sulfur
6.2 Classification of the amino acids by their frequency
Leu, Ala, Gly, Val, Glu, Ser, Ile, Lys, Arg, Asp, Thr, Pro, Asn, Gln,
Phe, Tyr, Met, His, Cys, Trp
7. MISCELLANEOUS STATISTICS
4473 entries are encoded on a mitochondrion, and 4063 are encoded on a plasmid.
12200 entries are encoded on a plastid,
of which 23 are encoded on apicoplasts,
11633 on chloroplasts,
51 on organellar chromatophores,
145 on cyanelles,
149 on non-photosynthetic plastids and
199 on unspecified types of plastid.
Number of entries with at least one sequence correction: 81583
Legend: gray = aliphatic, red = acidic, green = small hydroxy,
blue = basic, black = aromatic, white = amide, yellow = sulfur
6.2 Classification of the amino acids by their frequency
Leu, Ala, Gly, Val, Glu, Ser, Ile, Lys, Arg, Asp, Thr, Pro, Asn, Gln,
Phe, Tyr, Met, His, Cys, Trp
7. MISCELLANEOUS STATISTICS
4473 entries are encoded on a mitochondrion, and 4063 are encoded on a plasmid.
12200 entries are encoded on a plastid,
of which 23 are encoded on apicoplasts,
11633 on chloroplasts,
51 on organellar chromatophores,
145 on cyanelles,
149 on non-photosynthetic plastids and
199 on unspecified types of plastid.
Number of entries with at least one sequence correction: 81583
